Dysregulated expression of miRNAs can result disrupt proliferation, differentiation and apoptosis. Emerging evidence suggests that miRNAs are themselves regulated by endogenous molecules containing miRNA binding sites, such as pseudogenes, long noncoding RNAs, and circular RNAs. Termed 'miRNA sponges', these competitive inhibitors competitively sequester miRNAs, preventing them from interacting with their natural targets. The endogenous miRNA sponges and targets or competing endogenous RNAs (ceRNAs) act dynamically to balance the expression of each other. This balance is important for various physiological processes, and disruption can result in pathological states.
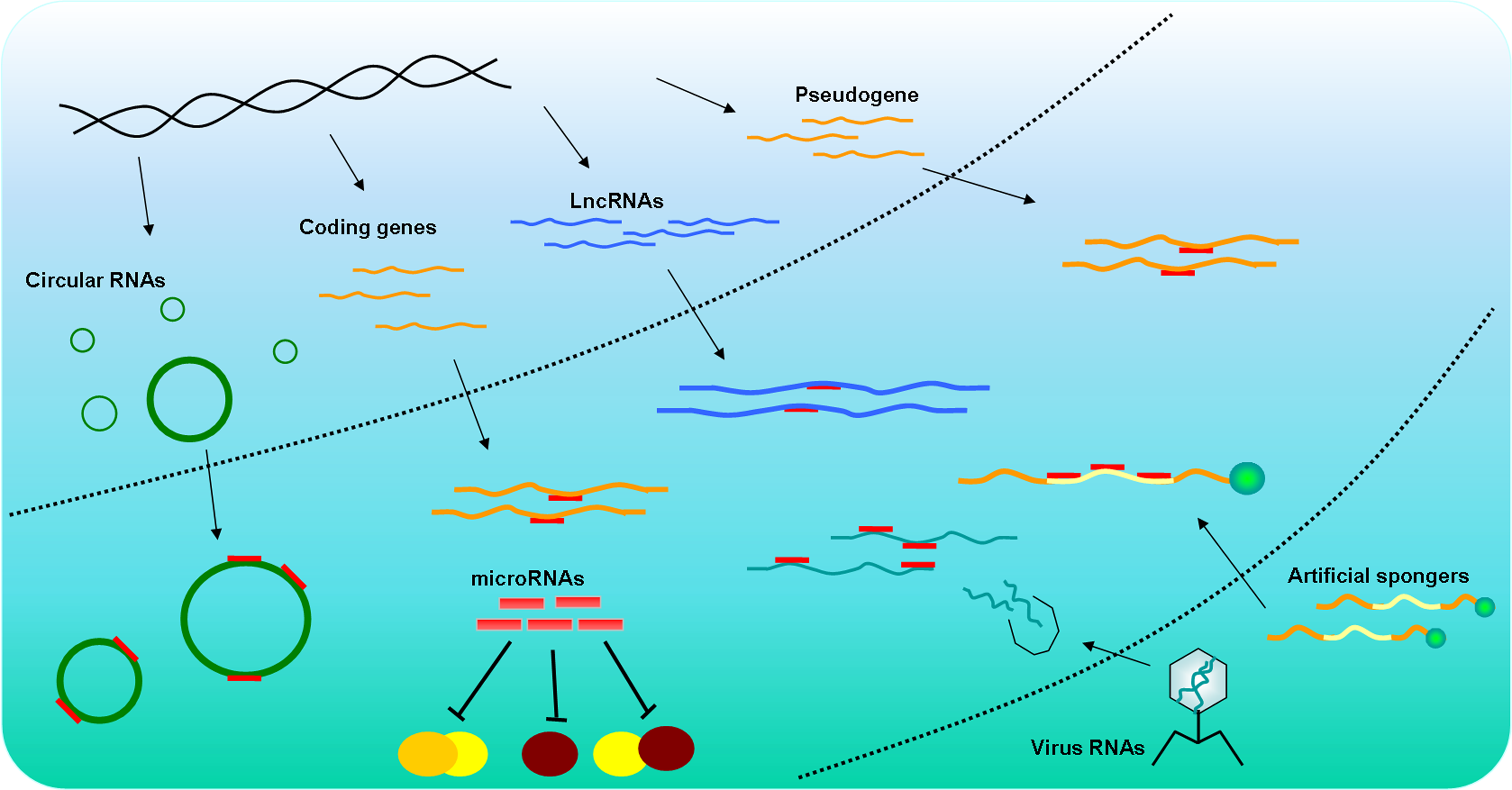
Statistics: The experimentally validated miRSponge database (miRSponge) recorded 576 sponge-miRNA interactions and 457 ceRNA associations in 11 species, following manual curating from nearly 1000 articles. Class of sponges include endogenously generated molecules such as coding-genes, pseudogenes, long noncoding RNAs and circular RNAs, and exogenously introduced molecules such as viral RNAs and artificial engineered sponges.
Around 70% of the interactions were experimentally identified in diseased states. Each entry contains detailed information on sponges, miRNAs and competing targets, species, miRBase accession, HGNC and GeneCards annotation, tissue distribution, conditions for detection, corresponding literature, PMIDs, a brief description of the sponge-miRNA interaction, and the experimental method used to characterize the association. miRSponge provides a user-friendly interface for convenient browsing, retrieval and downloading. In addition, miRSponge offers a submission page for newly validated miRNA-sponge and ceRNA association data.
In summary, the miRSponge database could significantly improve our understanding of transcriptome communication among different RNA classes, and provides a timely and valuable resource for miRNA research and clinical applications.