There is now overwhelming evidence that ncRNAs play pivotal roles in the regulation of immune system. First, ncRNAs regulate the innate and adaptive immune cell development and function. Furthermore, when ncRNAs are aberrantly expressed they can contribute to pathological conditions involving the immune system. In addition, ncRNAs can participate in the development of cancer by interacting with tumor immune microenvironment. Importantly, ncRNAs show great significance in modulating vaccine mediated innate and adaptive immune responses and disease prevention. To decode these series of regulation by ncRNAs, we describe a comprehensive database, RNA2Immune, which provides experimentally supported associations about ncRNA in immune regulation.
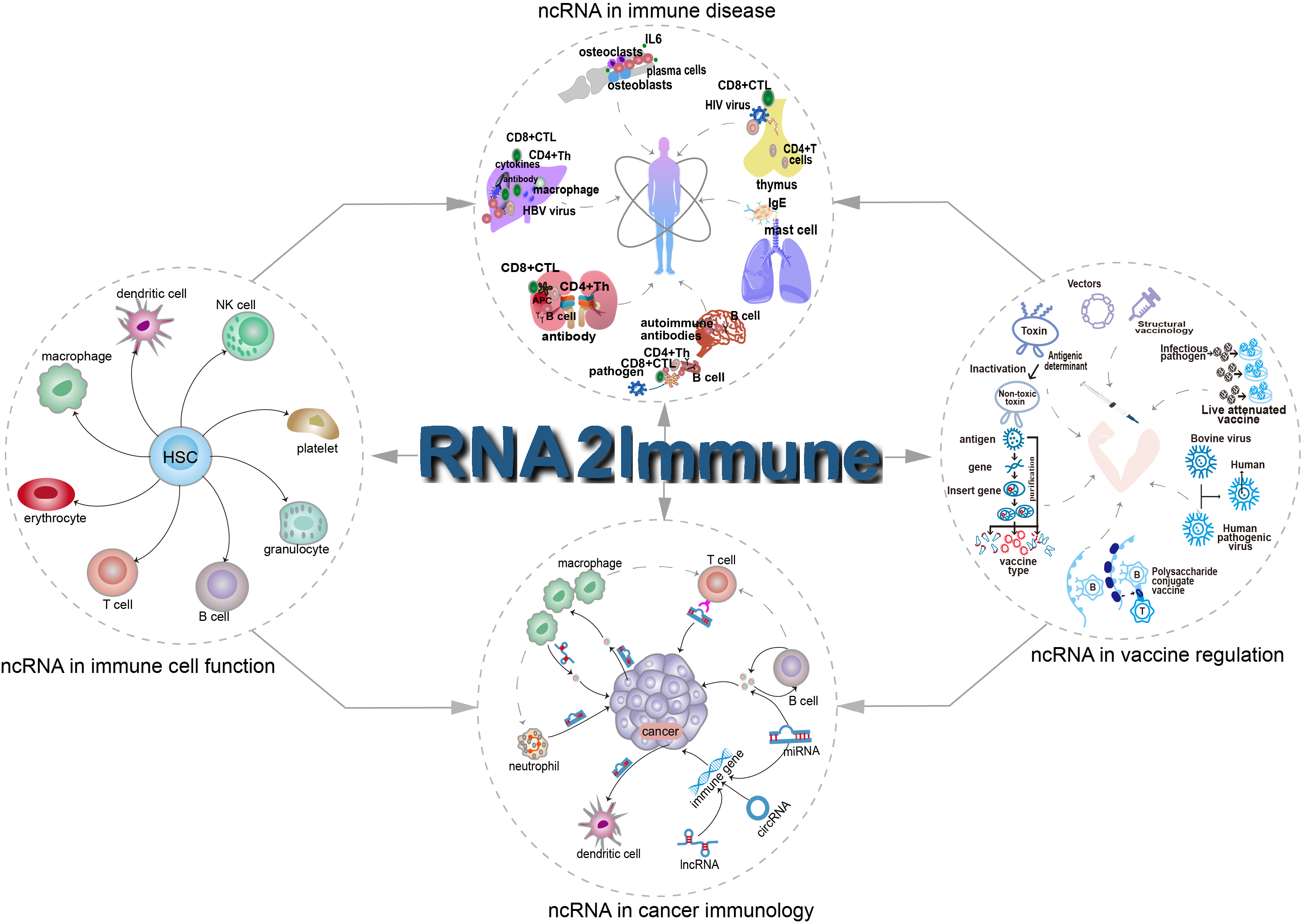
Figure 1 Overview of RNA2Immune
The current version of RNA2Immune documents 50433 associations about ncRNA in immune regulation in 42 host species, including: (i) 6690 ncRNA-influenced immune function entries from 31 immune cells; (ii) 38672 ncRNA-immune disease associations across 36 host species; (iii) 4833 ncRNA-immune cells/molecules mediated associations of cancers; (iv) 238 ncRNA-mediated associations of vaccine, referring to 26 vaccine types targeting 22 diseases. RNA2Immune provides a user-friendly searching, browsing and downloading interface. RNA2Immune provides important information about how ncRNA influence immune cells function, what pathological consequences (such as immune disease and cancer) of dysregulation of these ncRNAs, and how ncRNA modulate vaccine mediated disease prevention. Collectively, RNA2Immune will serve as an important resource for investigating the physiological and pathological mechanisms for ncRNAs in the immune system.
User can query the RNA2Immune database through Quick Search. Users can input an ncRNA or disease name or host species to search ncRNA-immune function associations, ncRNA-immune disease associations, ncRNA-immune cells/molecules mediated associations of cancers and ncRNA-vaccine associations. By clicking the "Number", a corresponding dataset will be displayed (as figure 2 below showing, taking the miRNA 'mmu-miR-155-5p' for example).
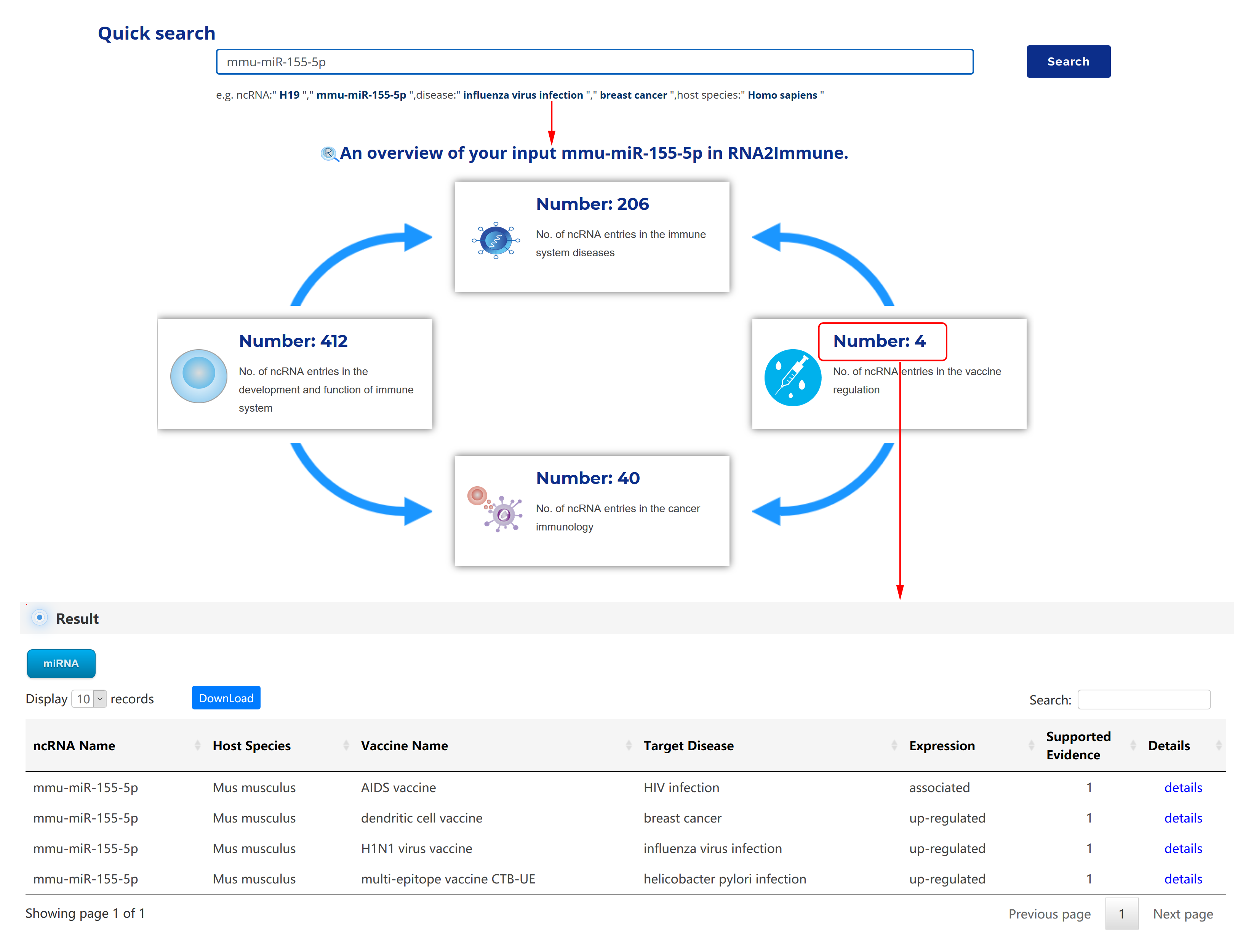
Figure 2 The use of quick search in the RNA2Immune
Immune function browse: To browse immune function part of the database, please click the menu ‘Browse’, then select ‘Immune Function’ (1 in figure 3 below). The browse page is classified by hierarchical piths of immune cells, ncRNA, immune function and host species. To browse the entries related a particular immune cell (for example CD8+ T cell), please click ‘Immune Cells’ (2, 3 in figure 3 below), then select ‘CD8+ T cell’ (4 in figure 3 below) in the catalog below. The result will be displayed on the right column (5 in figure 3 below). Various categories of ncRNA can be chosen in result column (6 in figure 3 below).
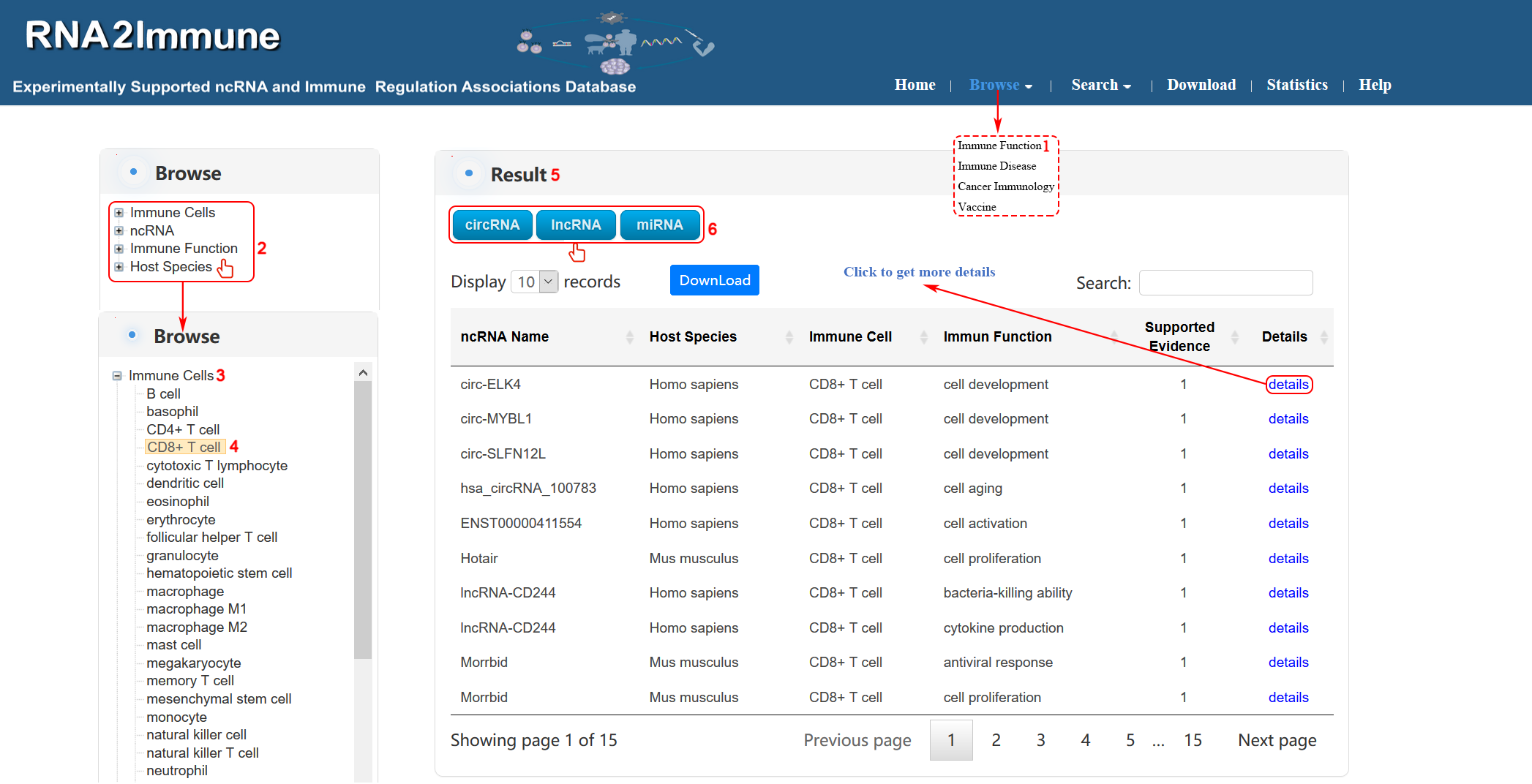
Figure 3 Interface of immune function browse
Immune Function search:To search Immune Function part of the database, please click the menu ‘Search’, then select ‘Immune Function’ (1 in figure 4 below).The Immune Function Search page provides Quick Search and Advanced Search. We have introduced Quick Search above. The Immune Function Advanced Search page allow user to search by combination of different key words. Users can select particular ncRNA Category from the pull-down menu (2 in figure 4 below). Input your interested ncRNA symbol in the 'ncRNA Name' search box (3 in Figure 4 below). Users can select all host species or particular species from the pull-down menu (4 in figure 4 below).Then, users can select the interested 'Immune cell' and 'Immune Function' in search box(5,6 in figure 4 below). At last, click the search button, the system will return to a table showing all entries (7,8 in figure 4 below).( As shown in figure 4 below, we take an example to explain).
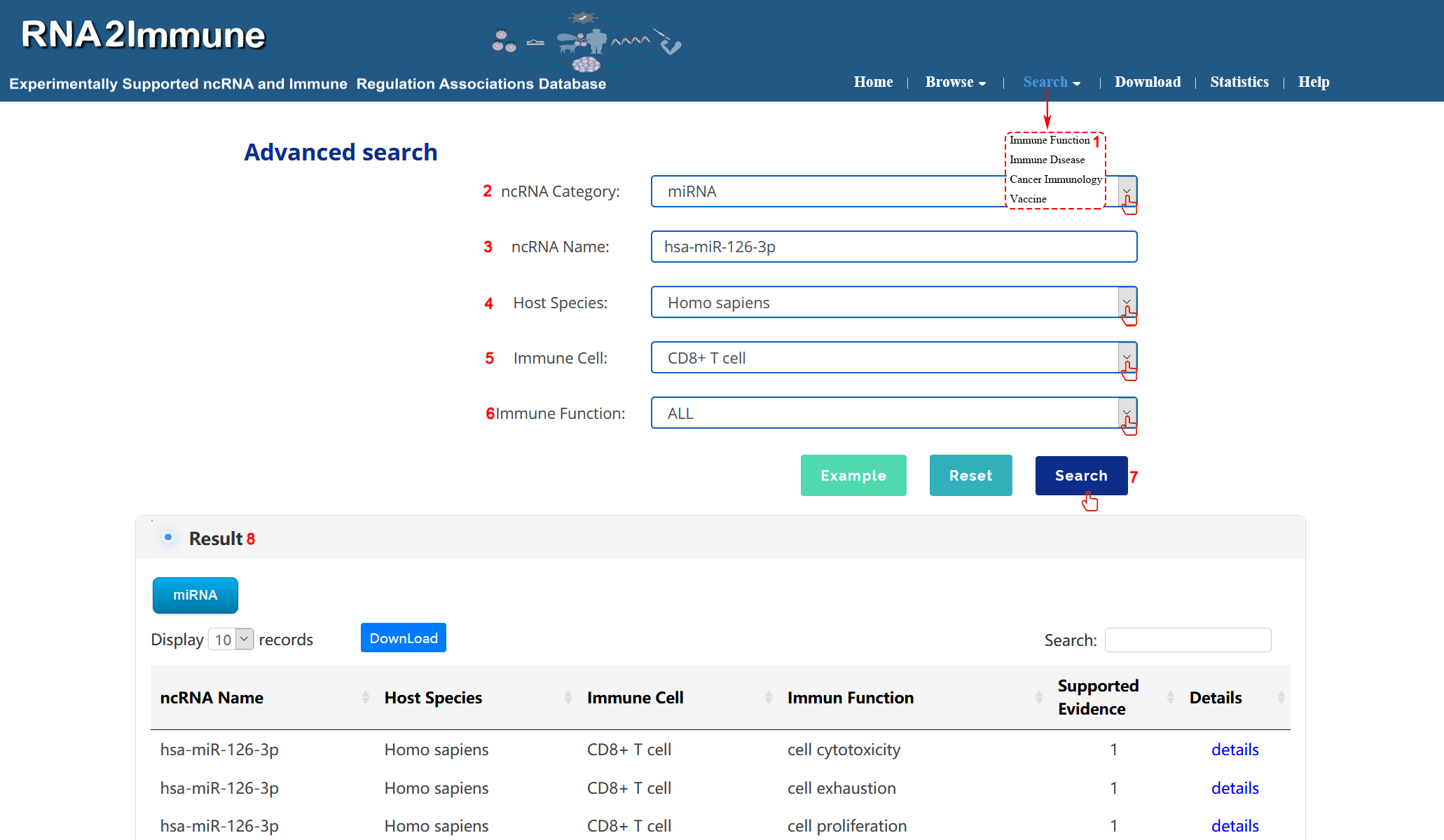
Figure 4 Interface of immune function search
Immune disease browse: To browse immune disease part of the database, please click the menu ‘Browse’, then select ‘Immune Disease’ (1 in figure 5 below). The browse page is classified by hierarchical piths of immune disease, ncRNA and host species (2 in figure 5 below). To browse the entries related a particular immune disease name(for example avian leukosis), please click ‘Immune Disease’(3 in figure 5 below), then choose the subcategory ‘Infectious Diseases’, and select ‘Virus Infection’ which is corresponding disease ‘avian leukosis’(4 in figure 5 below) in the catalog below. The result will be displayed on the right column (5 in figure 5 below). Various categories of ncRNA can be chosen in result column (6 in figure 5 below).
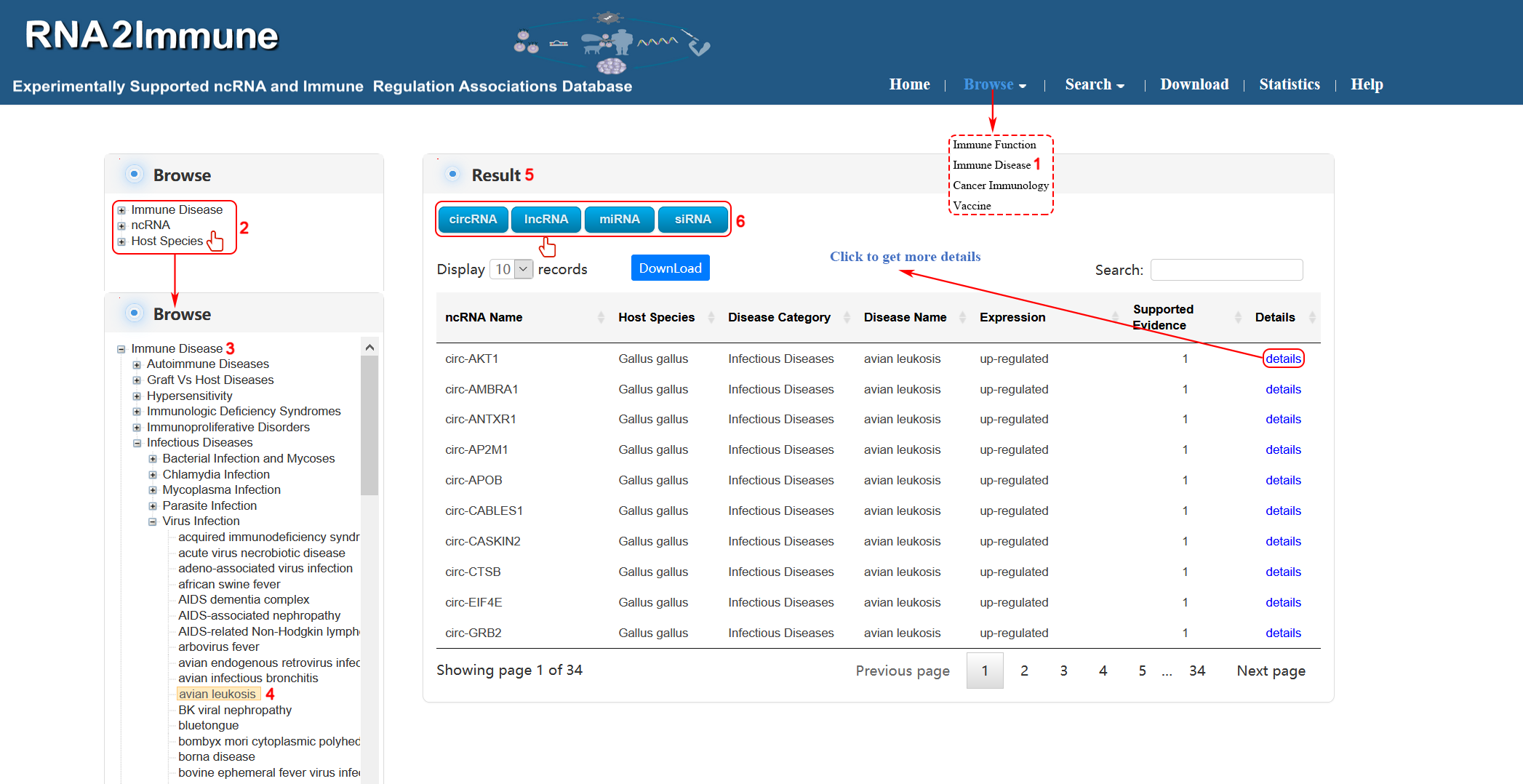
Figure 5 Interface of immune disease browse
Immune Disease search:To search immune disease part of the database, please click the menu ‘Search’, then select ‘Immune Disease’(1 in figure 6 below).Users can select particular ncRNA category in search box (2 in figure 6 below). Input your interested ncRNA symbol in the 'ncRNA Name' search box (3 in Figure 6 below). Then choose all host species or particular species from the pull-down menu (4 in figure 6 below). Select all disease or your interested disease category in the 'Disease Category' search box (5 in figure 6 below). Choose your interested disease in the 'Disease Name' search box (6 in figure 6 below). At last, click the search button, the system will return to a table showing all entries (7,8 in figure 6 below). (As shown in figure 6 below, we take an example to explain).
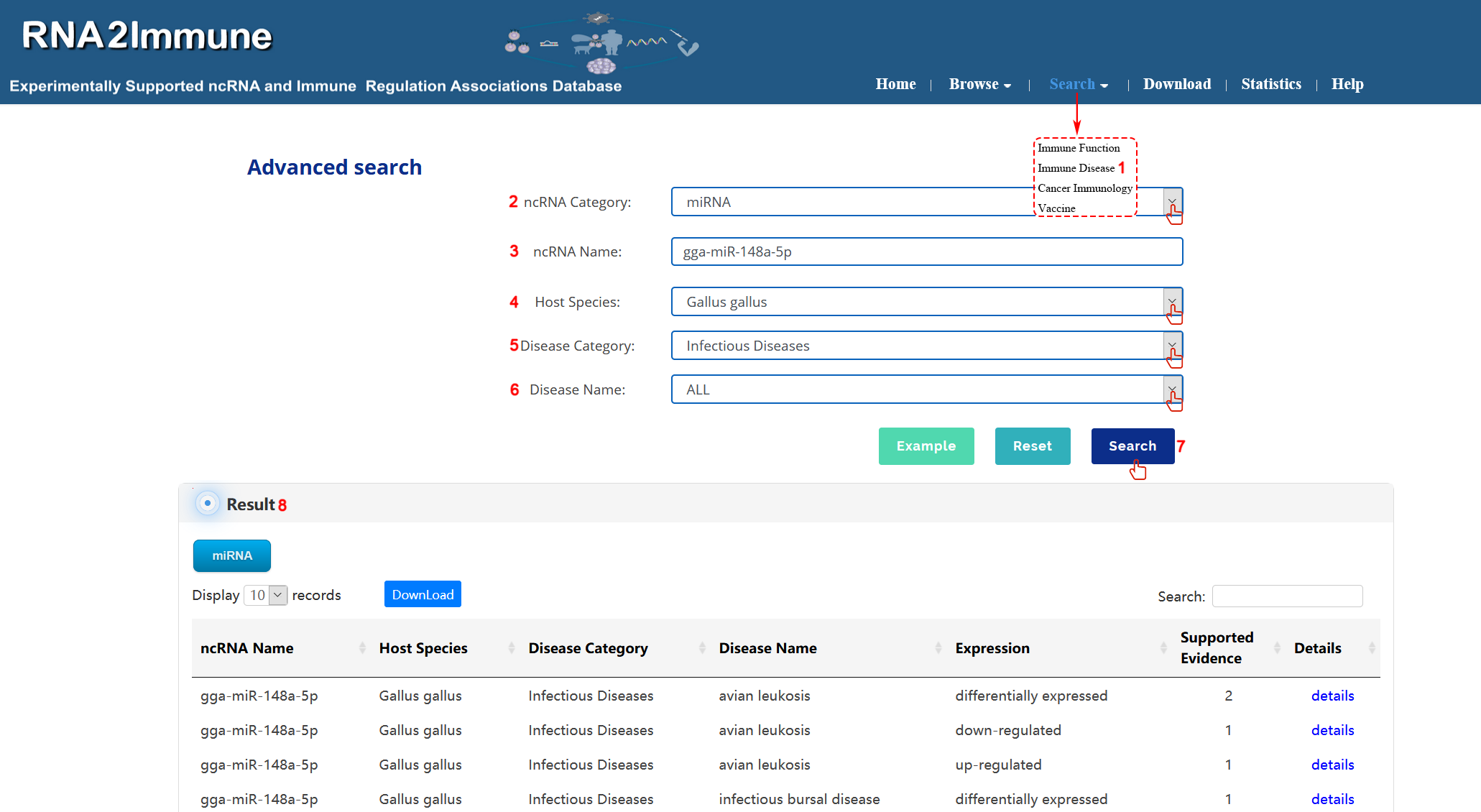
Figure 6 Interface of immune disease search
Cancer immunology browse: To browse cancer immunology part of the database, please click the menu ‘Browse’, then select ‘Cancer Immunology’ (1 in figure 7 below). The browse page is classified by hierarchical piths of cancer, ncRNA, immune cells, immune molecules and host species (2 in figure 7 below). To browse the entries related a particular cancer (for example breast cancer), please click ‘cancer’ (3 in figure 7 below), then select ‘breast cancer’ (4 in figure 7 below) in the catalog below. The result will be displayed on the right column (5 in figure 7 below).Various categories of ncRNA can be chosen in result column (6 in figure 7 below).
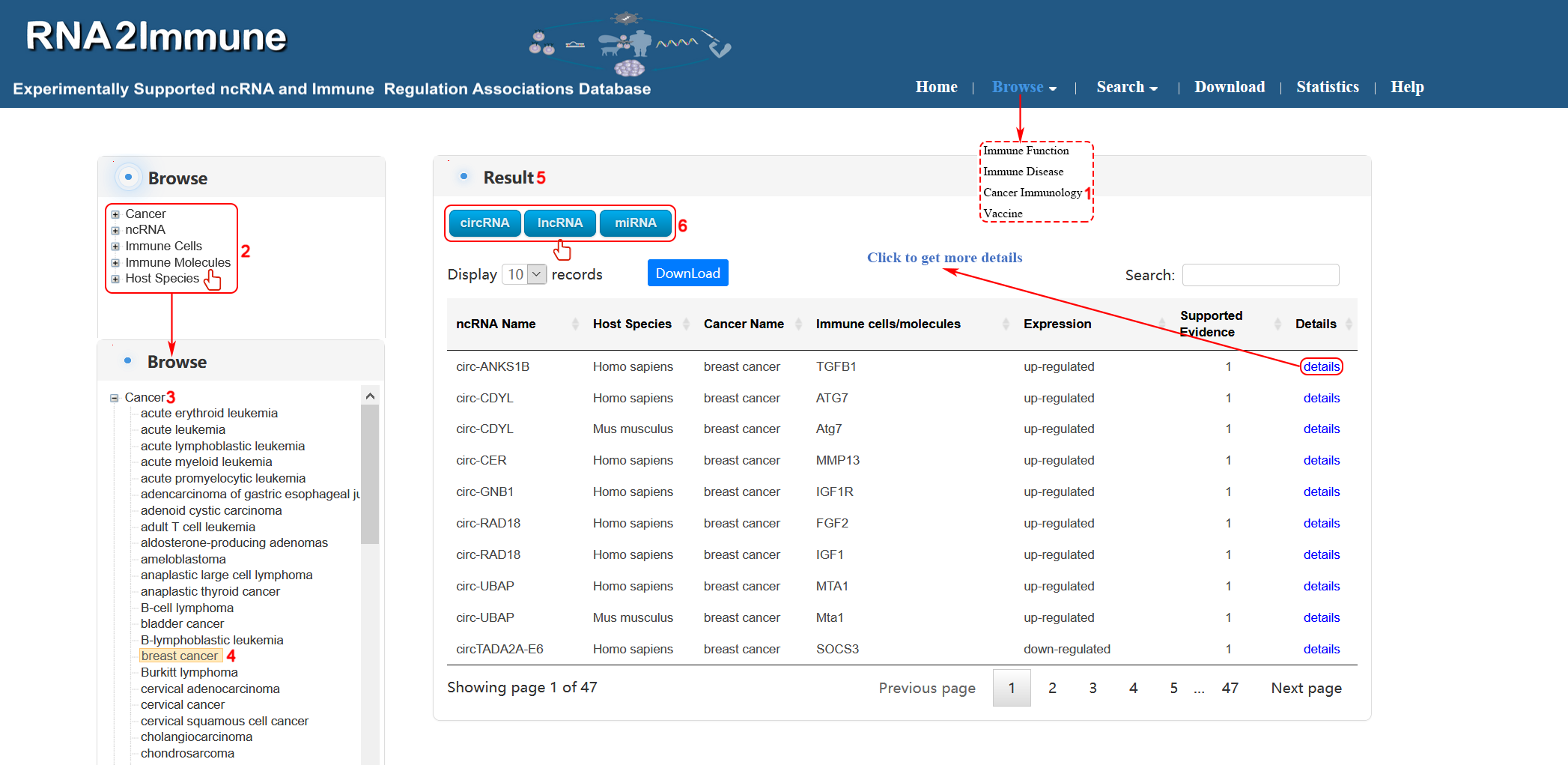
Figure 7 Interface of cancer immunology browse
Cancer immunology search: To search cancer immunology part of the database, please click the menu ‘Search’, then select ‘Cancer Immunology’ (1 in figure 8 below).Users can select particular ncRNA category in search box (2 in figure 8 below). Input your interested ncRNA symbol in the 'ncRNA Name' search box (3 in Figure 8 below). Then choose all host species or particular species from the pull-down menu (4 in figure 8 below). Select all cancer or your interested cancer name in the 'Cancer Name' search box (5 in figure 8 below). Choose your interested Immune cells or molecules in filter box (6,7 in figure 8 below). At last, click the search button, the system will return to a table showing all entries (8,9 in figure 8 below). (As shown in figure 8 below, we take an example to explain).
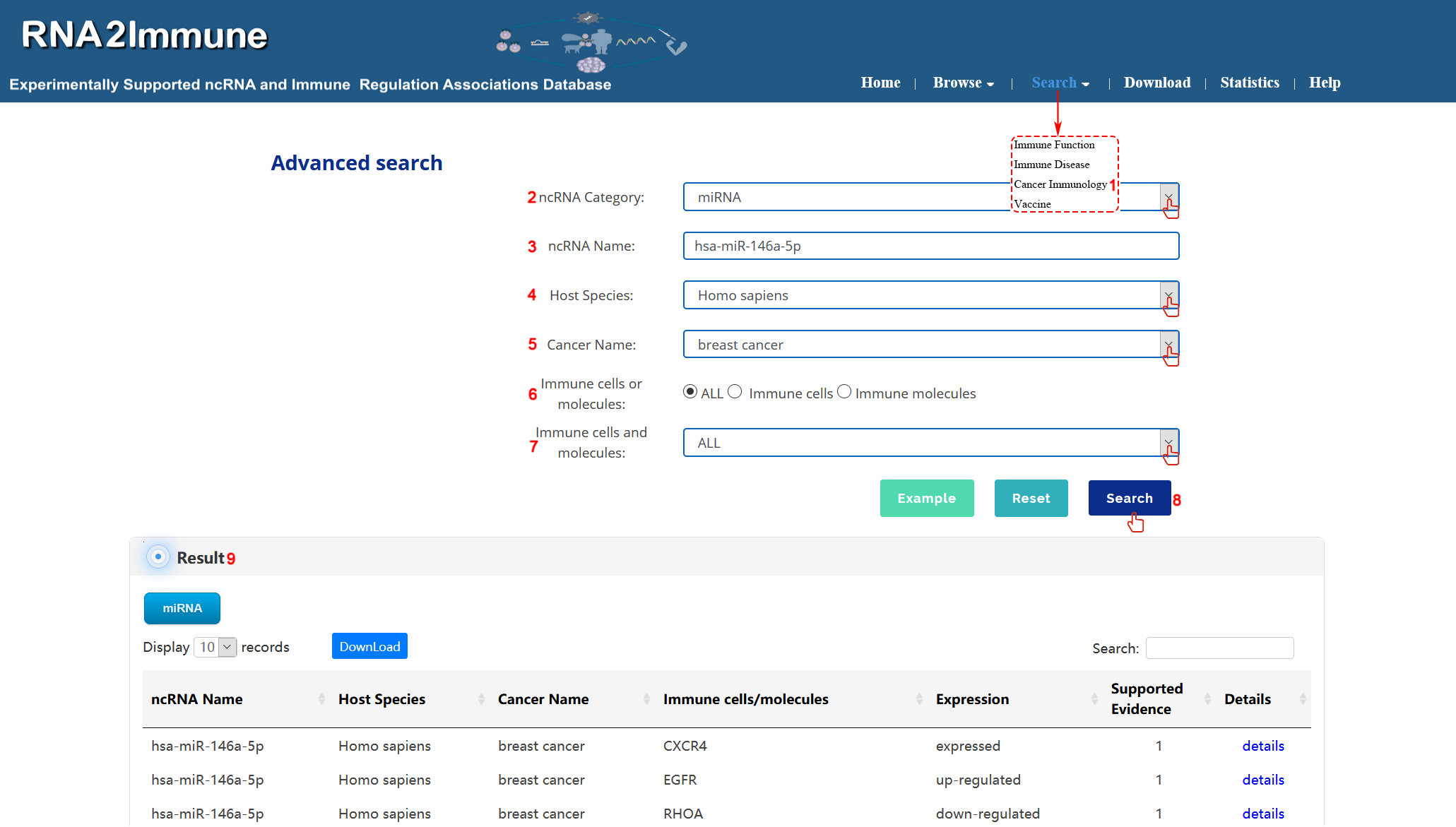
Figure 8 Interface of cancer immunology search
Vaccine browse: To browse vaccine part of the database, please click the menu ‘Browse’, then select ‘Vaccine’ (1 in figure 9 below). The browse page is classified by hierarchical piths of vaccine name, ncRNA, target disease and host species (2 in figure 8 below). To browse the entries related a particular vaccine name (for example Bacillus Calmette-Guerin), please click ‘Vaccine Name’ (3 in figure 9 below), then select ‘Bacillus Calmette-Guerin’ in the ‘live attenuated vaccine’ catalog below (4 in figure 9 below). The result will be displayed on the right column (5 in figure 9 below).Various categories of ncRNA can be chosen in result column (6 in figure 9 below).
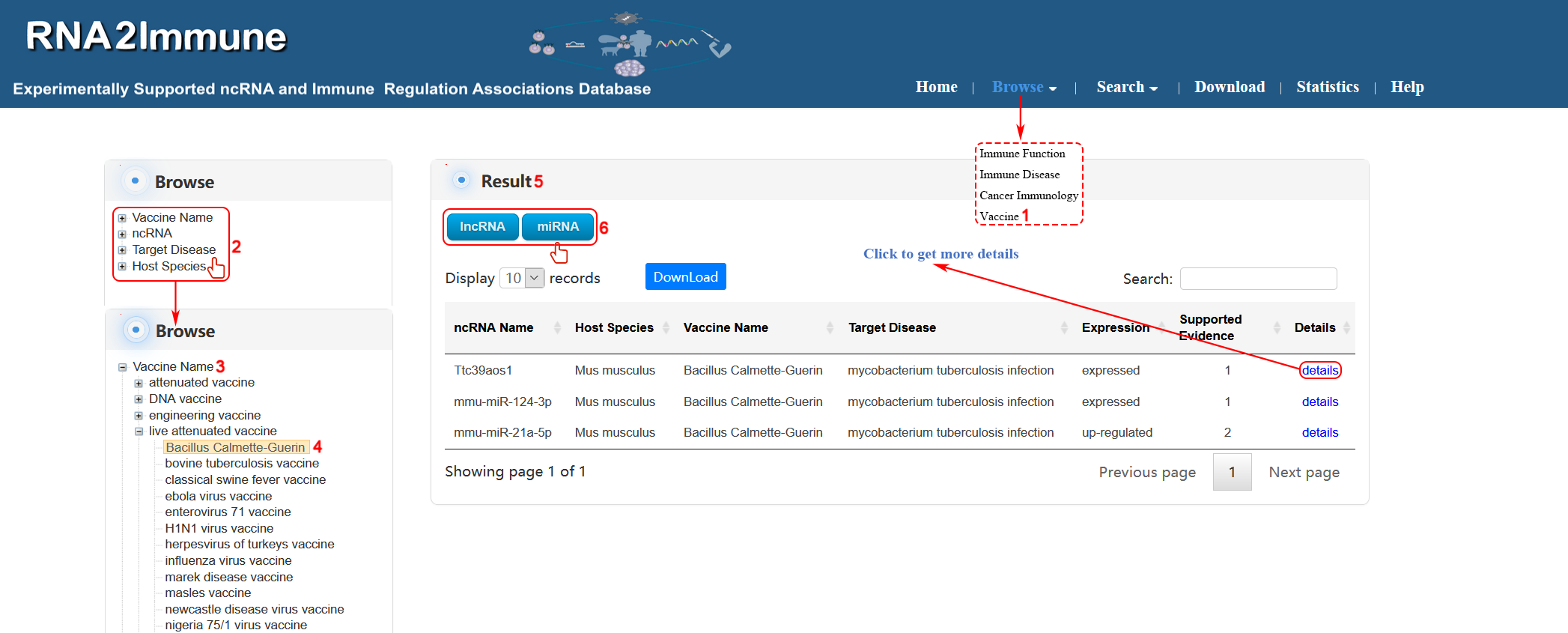
Figure 9 Interface of vaccine browse
Vaccine search: To search vaccine part of the database, please click the menu ‘Search’, then select ‘Vaccine’ (1 in figure 10 below). Users can select particular ncRNA category in search box (2 in figure 10 below). Input your interested ncRNA symbol in the 'ncRNA Name' search box (3 in Figure 10 below). Then choose all vaccine name or particular vaccine from the pull-down menu (4 in figure 10 below). Select all target disease or your interested disease in the search box (5 in figure 10 below). Choose all host species or particular species from the pull-down menu (6 in figure 10 below). At last, click the search button, the system will return to a table showing all entries (7,8 in figure 10 below). (As shown in figure 10 below, we take an example to explain).
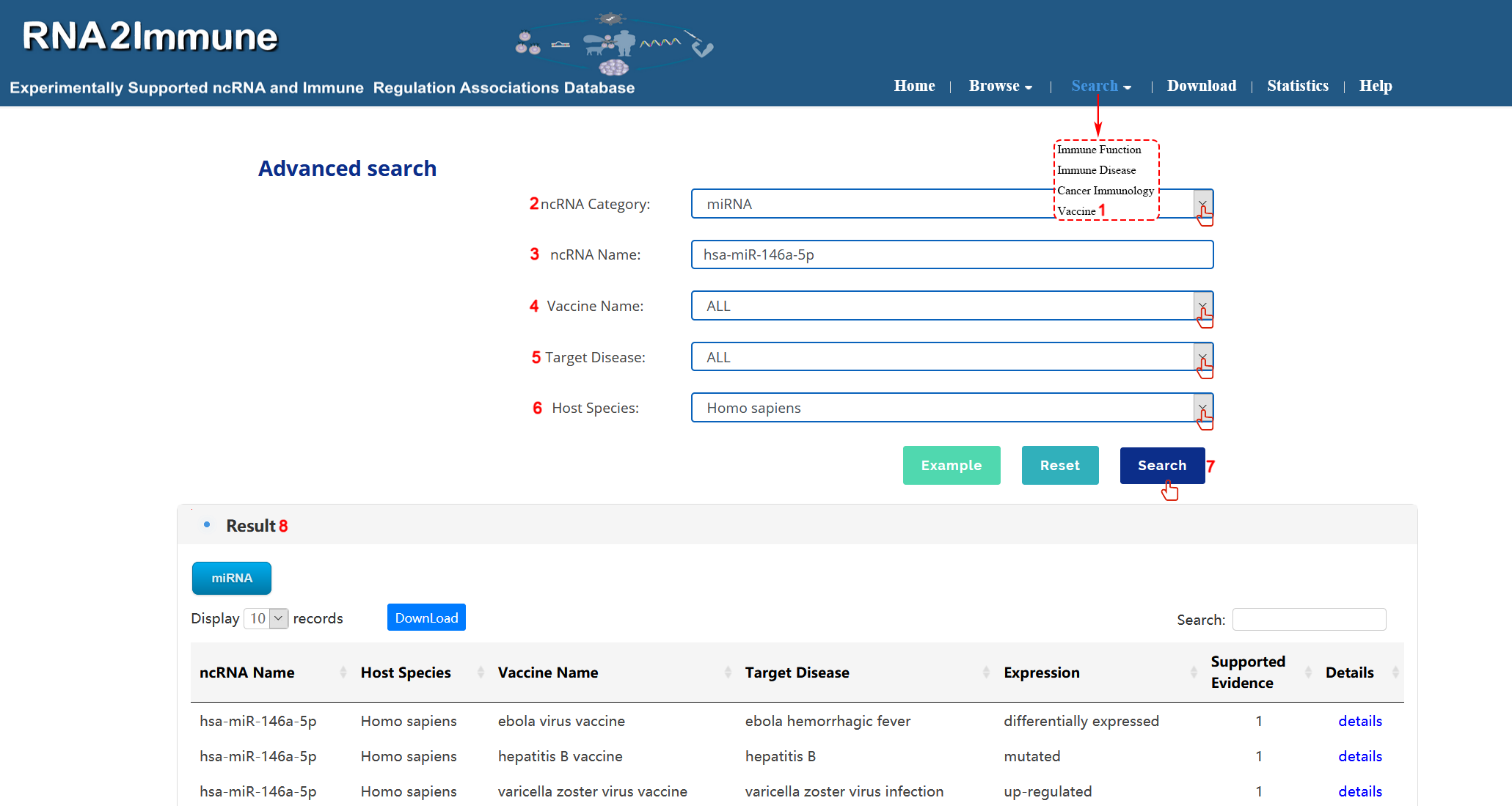
Figure 10 Interface of vaccine search
Users can obtain detailed information from each supported ncRNA by clicking the 'more details' of the data table. In the detail page, users can obtain basic information for the ncRNA (1 in figure 11 below). Then, users can search the external annotation information for the ncRNA by clicking on the corresponding hyperlink (2 in figure 11 below). Moreover, users can obtain detailed information for the ncRNA and Immune Function, Immune Disease, Cancer Immunology or Vaccine (3,4 in figure 11 below). Finally, users can get information of each ncRNA from various supporting evidence (5 in figure 11 below).
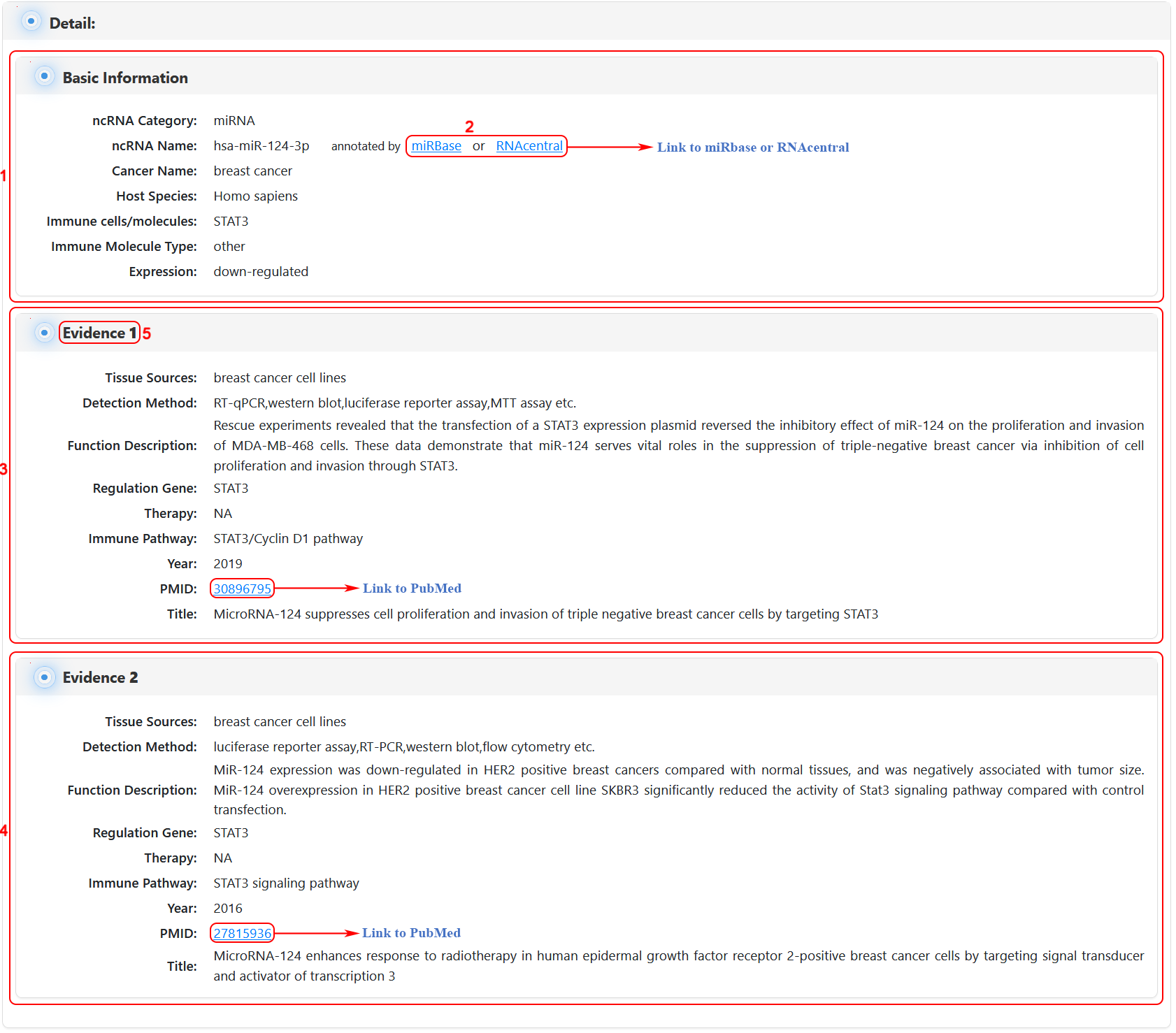
Figure 11 Interface of detailed information
To download data in the database, select the menu 'Download'. RNA2Immune database provides downloadable file in TXT format. Users can download all data sets, which include four files: (i) All experimentally supported ncRNA-immune function associations of different immune cell types across 15 host species; (ii) All experimentally supported ncRNA-immune disease associations across 36 host species; (iii) All experimentally supported ncRNA-immune cells/molecules mediated associations of cancers; (iv) All experimentally supported ncRNA-vaccine associations across 9 host species (figure 12).
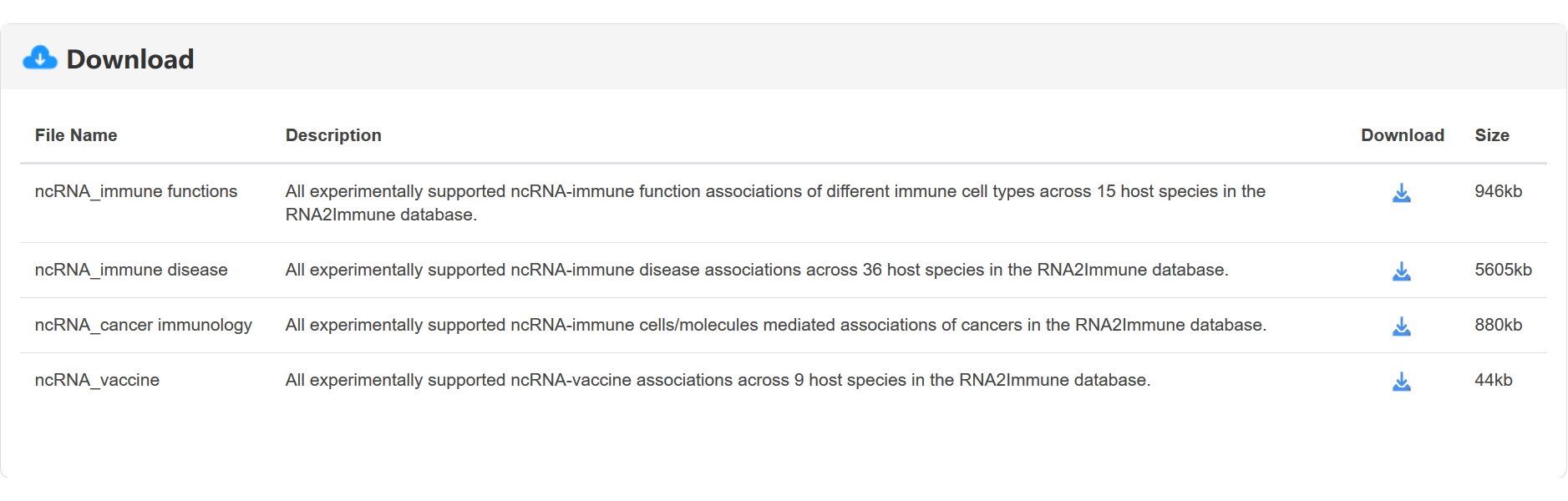

Figure 12 Interface of the download module
Please contact us when you have any questions in using this website.
Address
194 Xuefu Road, Harbin 150081, CHINA
Lihua Wang:wanglh211@163.com or Shangwei Ning:ningsw@ems.hrbmu.edu.cn