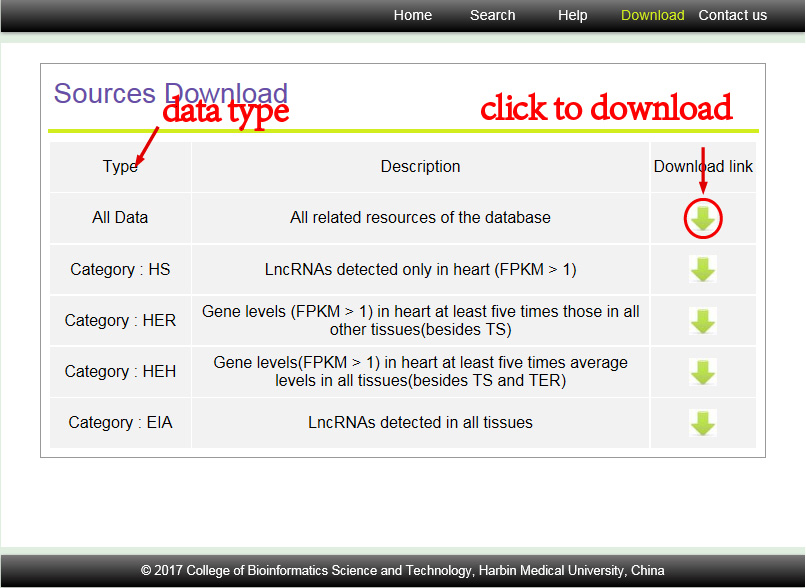
Long noncoding RNAs (lncRNAs) have been revealed to play essential roles in the human cardiovascular system. However, information about their mechanisms is limited, and a comprehensive view of cardiac lncRNAs is lacking from a multiple tissues perspective to date. Here, the landscape of the lncRNA transcriptome in human heart was summarized. We summarized all lncRNA transcripts from publicly available human transcriptome resources (156 heart samples and 210 samples from 29 other tissues) and systematically analysed all annotated and novel lncRNAs expressed in heart. A total of 7,485 lncRNAs whose expression was elevated in heart (HE lncRNAs) and 453 lncRNAs expressed in all 30 analysed tissues (EIA lncRNAs) were extracted. Using various bioinformatics resources, methods and tools, the features of these lncRNAs were discussed from various perspectives, including genomic structure, conservation, dynamic variation during heart development, cis-regulation, differential expression in cardiovascular diseases and cancers, and regulation at transcriptional and post-transcriptional levels. Afterwards, all the features discussed above were integrated into a user-friendly resource named CARDIO-LNCRNAS. This study represents the first global view of lncRNAs in the human cardiovascular system based on multiple tissues and sheds light on the role of lncRNAs in developments and heart disorders.
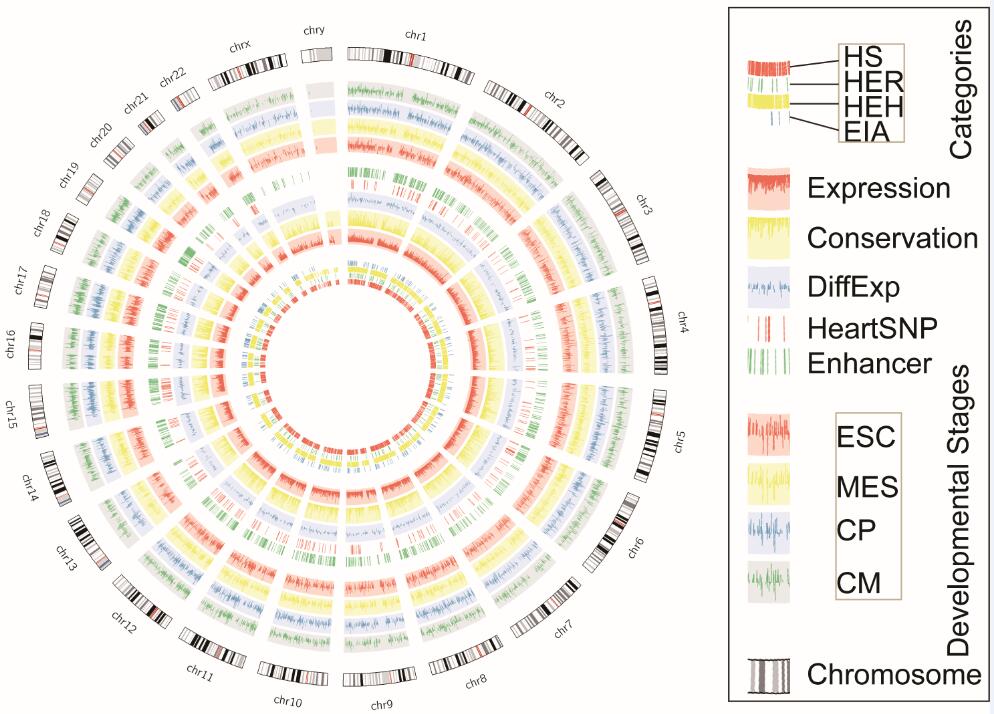
HS, HER, and HEH are the three subtypes of HE lncRNAs, whose expressions are elevated in human heart tissue. EIA are lncRNAs expressed in all tissues analysed. Expression represents log10FPKM of each lncRNA in human heart tissue. Conservation represents the PhastCons scores of exons for each lncRNA. DiffExp represents the -log10P, where the P is the significance of differential expression in cardiovscular diseases. HeartSNP is the heart related SNPs lacated around 100kb of a lncRNA. Enhancer is the enhacners from seven heart cell lines of FANTOM5 project, who locate in 1kb to 50kb around a lncRNA. ESC, MES, CP, CM are the expression of lncRNAs in four developmental stages of heart tissue.
Two ways were provided to scan the information of your interested lncRNA or lncRNA set in detail.
If you are interested in a chromosome region, and you want to see the information of all lncRNAs in this region, you can click the region in the chromosome of the Circos atlas in the home page.
Step 1 : Choose a chromosome you interested, click the chromosome to amplify it.
Step 2 : Choose the detailed area you want to view, click the area to search
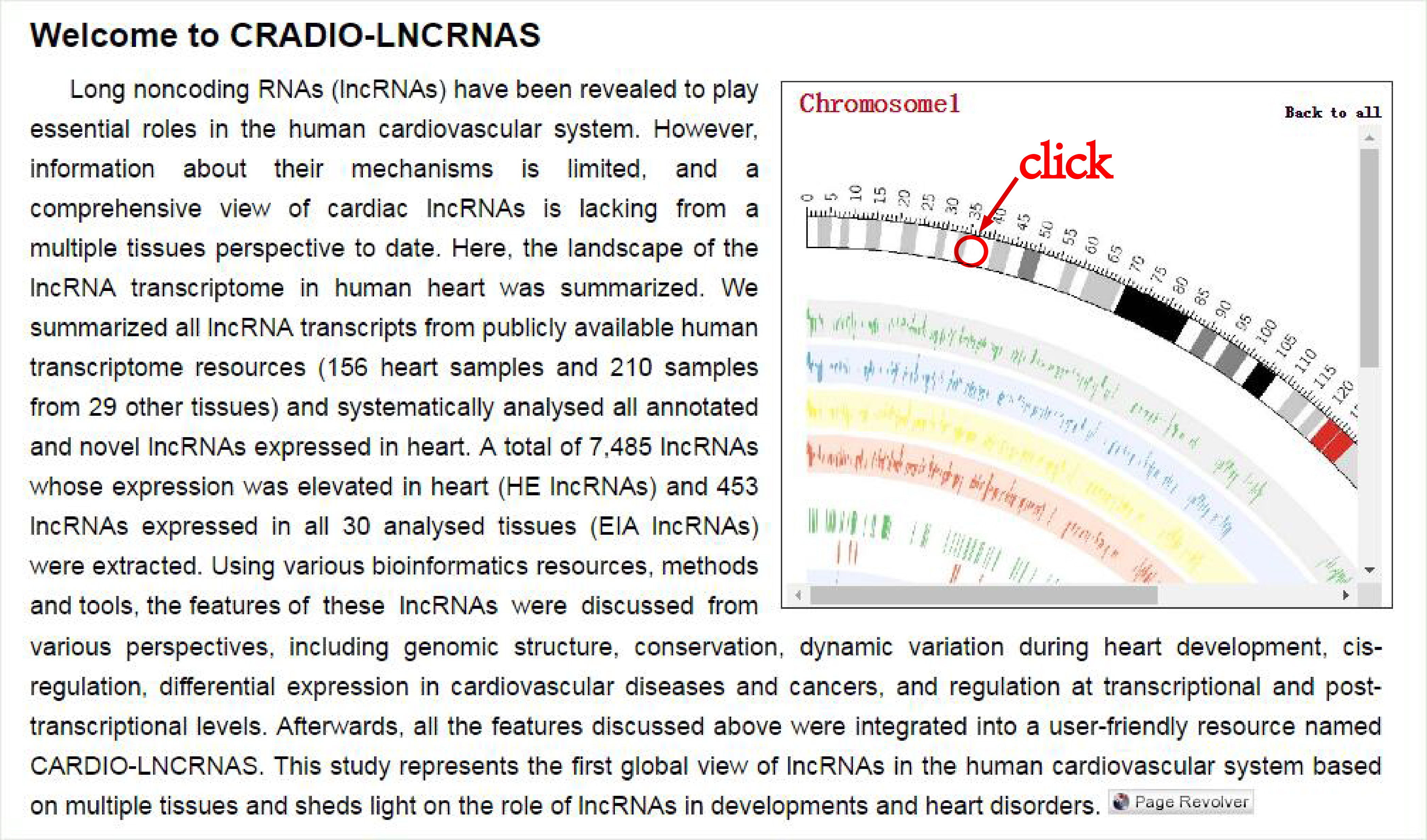

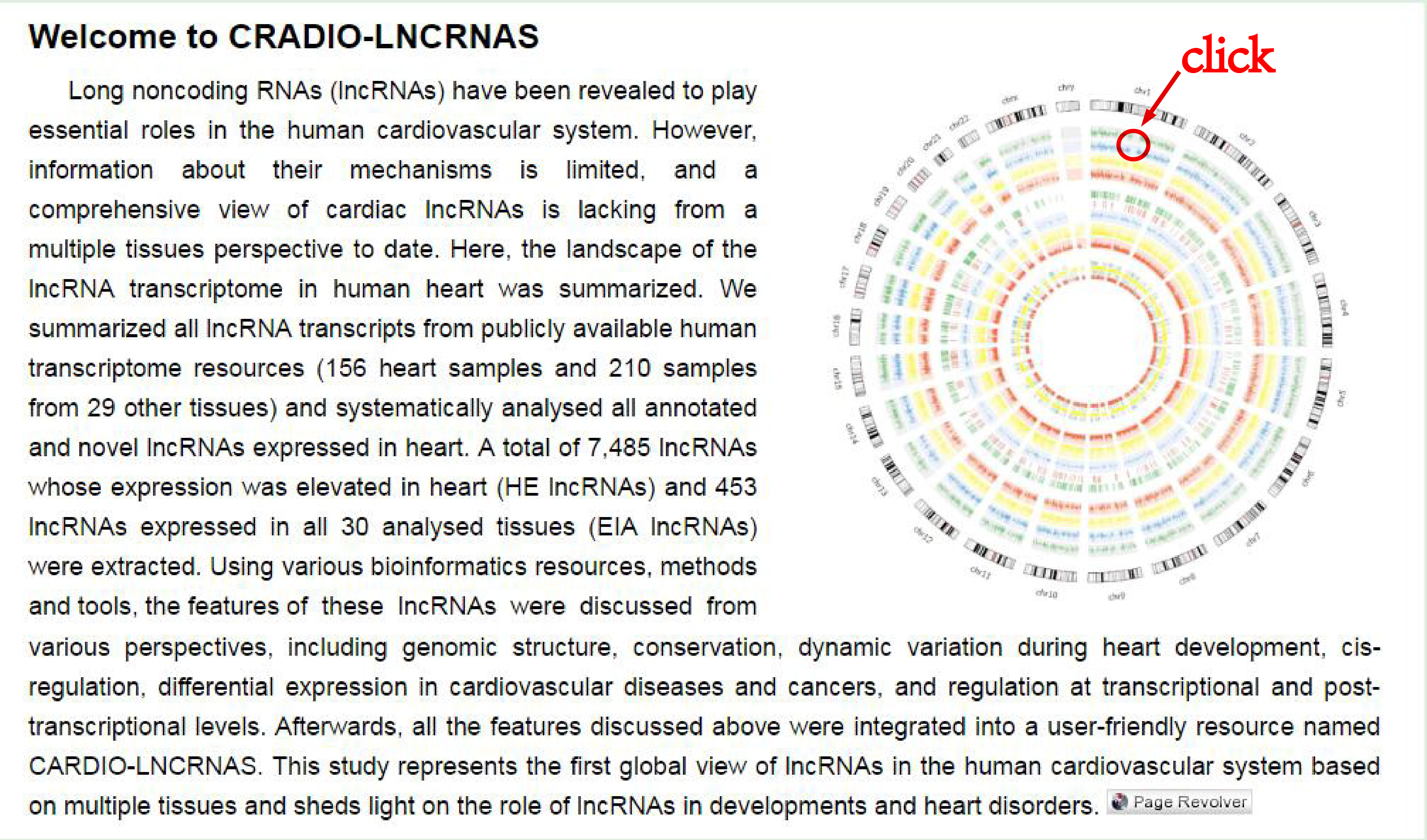
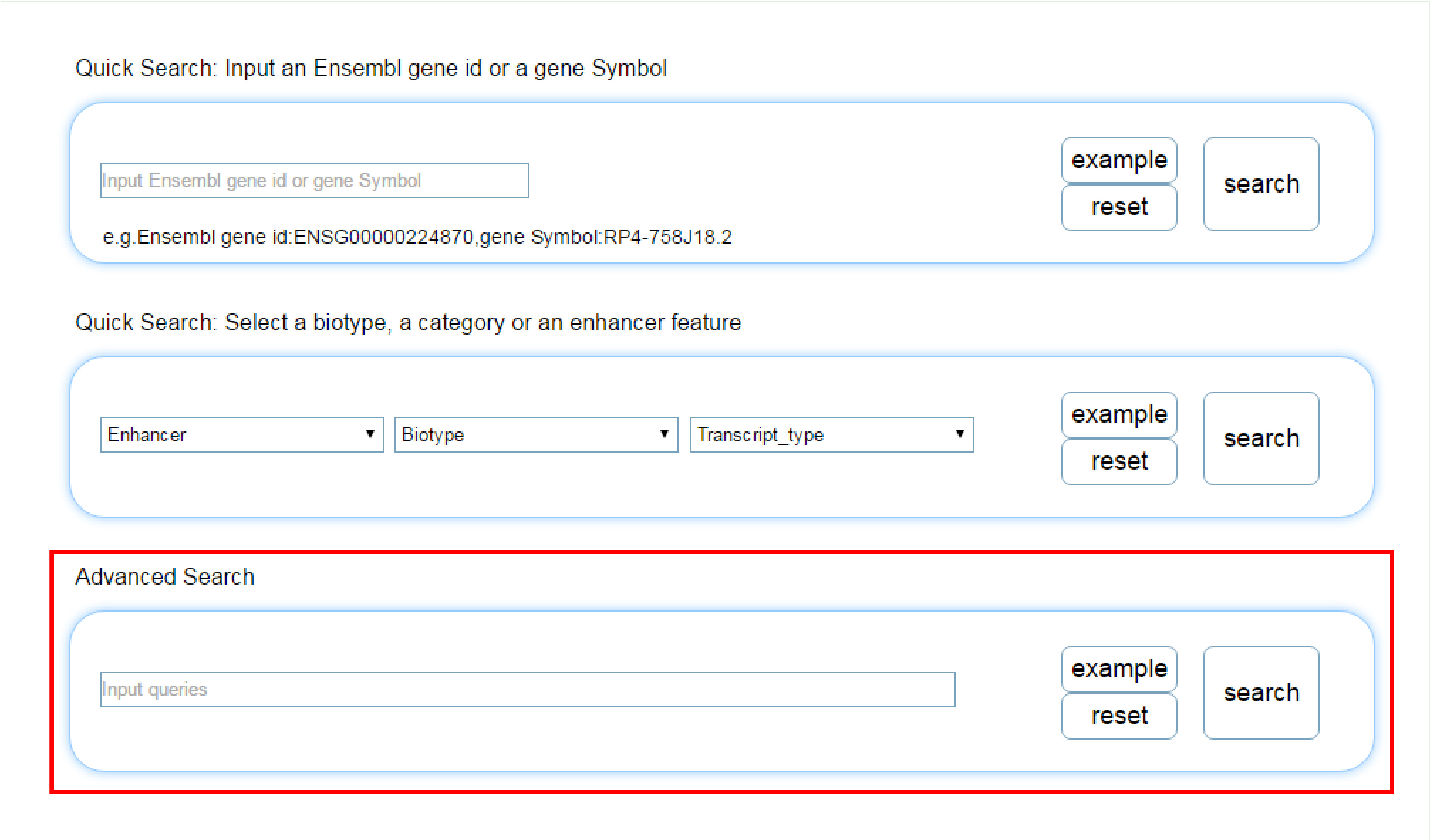
You can also view the information of the lncRNA through inputing Ensembl Id or Symbol, or by selecting biotype, category, or enhancer in the searching page.
The Advanced search allows for greater control of search results by allowing queries based on various types of data included in CARDIO-LNCRNAS. The advance search uses a query tag combined with a search string to specify the type of data to return. Additionally, the output of these queries can be combined together using the boolean operators: “and", “or", “not", and “()”. Note, if a query contains a space, then quotation marks should be added around the full string. If no tag is present in the query, it will be treated as a biotype. Again, if any spaces are included within the string, the entire string will be treated as multiple queries unless surrounded by quotation marks. See the table below for the types of queries:
| Expression | Example | Description |
|---|---|---|
| No tag | Intergenic | When no tag is specified the query is treated as a biotype. |
| Biotype | Intergenic | Returns lncRNAs that biotype is Intergenic. |
| Category | EIA | Returns lncRNAs that category is EIA. |
| Enhancer | YES | Returns lncRNAs that enhancer is YES. |
| Boolean Operators | ||
| and | Biotype:Antisense and Biotype:Intergenic | Returns accessions existing between two query sets. |
| or | Biotype:Antisense or Biotype:Intergenic | Returns accessions existing in either of two query sets. |
| not | not Biotype:Intergenic | Returns accessions not in the query. |
| () | Biotype:Antisense or (Biotype:Intergenic not Category:EIA and Enhancer:YES) | Increases the precedence of a query. |
| * | Interge* | Specifies where to use wildcards in query. Note: wildcards are only supported for tagless searches. |
You can get the result through the two ways described above.
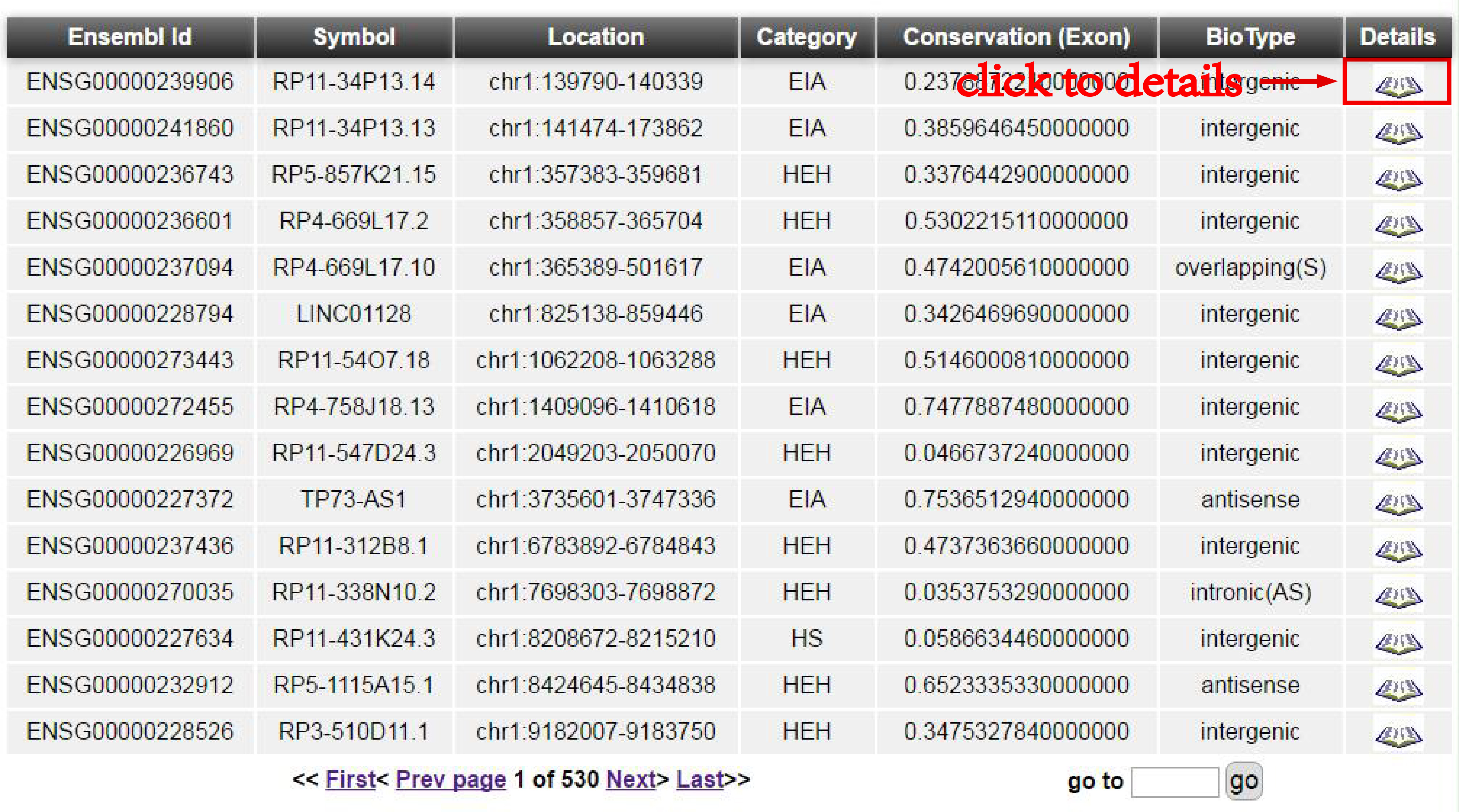
More importantly, you can see more information of a selected lncRNA through the detail link in the result page. And you can visualize the expression in each heart sample, each tissue and each developmental sample.
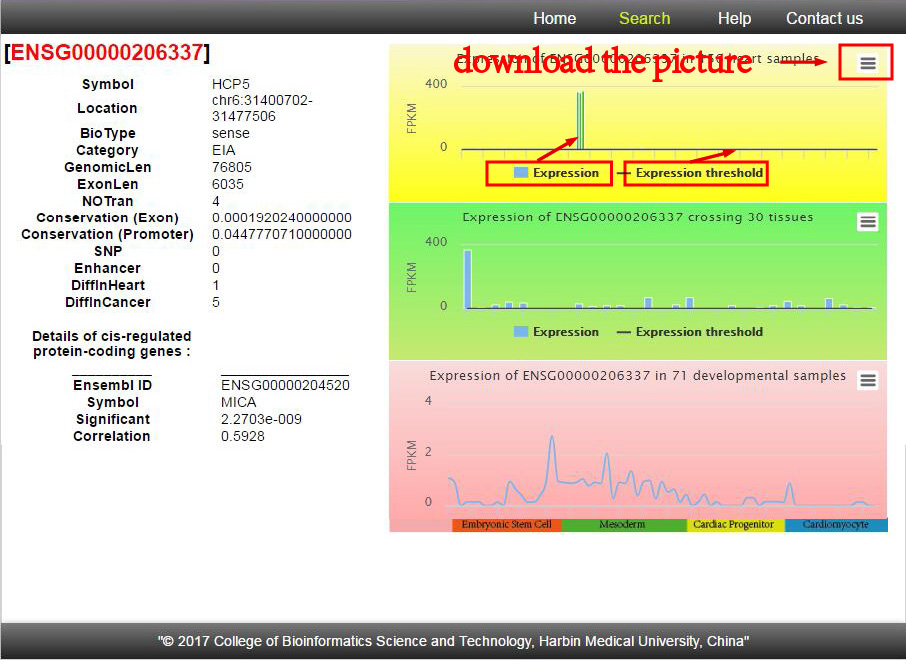
In addition, download links are provided, which allowed you to batch download the information you interested in.