Help |
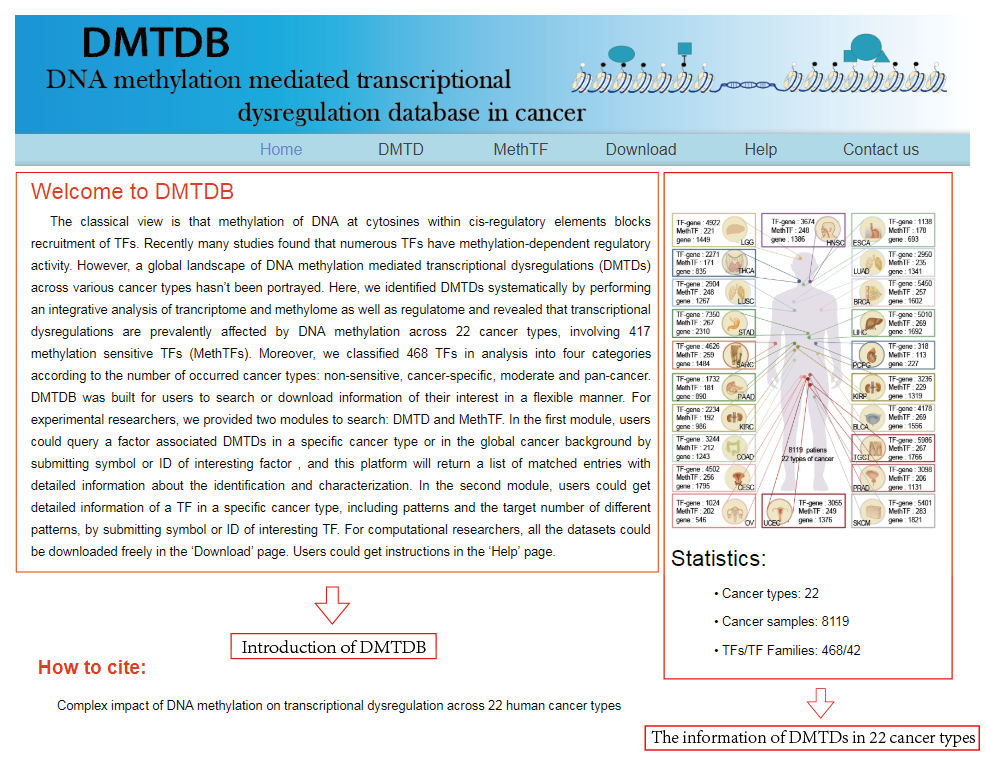
Welcome to DMTDB! Here is a step to step introduction about how to use the web to search the DMTDs and MethTFs identified in 20 cancer types.

Here, we used a three-step computational framework to identify DMTDs in a specific cancer type.
Step-1,from ChIPBase v2.0 we downloaded ChIP-seq peak regions of 468 TFs, which were intersected with promoter regions of genes to identify genome-wide TF-gene regulations.
Step-2, we identified cancer-context TF-gene regulations in each cancer type. We used linear regression to measure context-dependent TF-gene expression association in different cancer types. In this model, mRNA i expression(log2), yi, changes as a linear function of TF μ expression(log2),xμ,among the n cancer samples of a given cancer type: yi=β0+βμxμ,i+εi,i=1,…,n where β0 is the intercept, βμ is the regression coefficients for TF expression variable. We estimated the TF coefficient by using ordinary least squares method to test whether the expression level of gene is associated with changes of expression level of TF. TF-gene regulations with Benjamini and Hochberg (BH) adjusted P-value less than 0.01 were regarded as cancer-context TF-gene regulations for further analysis.
Step-3, we identified DMTDs from cancer-context TF-gene dysregulations in each cancer type based on the changes of TF’s regulatory activity mediated by DNA methylation. TFs and target genes were filtered based on the expression variation across samples. Individual TFs and target genes with highly variation across samples were selected for further analysis (log2 IQR > 0.58). For each cancer-context TF-gene dysregulation, the cancer samples were sorted based on the methylation level of target gene and we defined the top and bottom 25% samples as high-methylation group (H-group) and low-methylation group (L-group). For each cancer type, we calculated the difference of average methylation level of target genes between H-group and L-group for all cancer-context TF-gene regulations. We only analyzed the top 25% TF-gene dysregulations in terms of the difference of average methylation level of target genes between H-group and L-group. In addition, a TF-gene dysregulation must fulfill the following two criteria for further analysis: i) the TF wasn’t differentially expressed between H-group and L-group; ii) the target gene was differentially expressed between H-group and L-group. Next, we tested each TF-gene dysregulation to determine whether it was affected by DNA methylation. We used the Spearman correlation coefficient between expression of TF and gene from a TF-gene dysregulation to measure the regulatory activity of the TF in the H-group (Rhigh) and L-group (Rlow), separately. To guarantee that TF regulated gene in at least one condition, we required that the absolute value of either Rhigh or Rlow was > 0.3. Then we selected TF-gene dysregulations with the absolute value of difference between Rhigh and Rlow > 0.3 for further analysis. Next, Fisher’s test of difference between two Spearman correlation coefficients was used. Briefly, we transformed these Spearman correlation coefficients between TFs and genes as follow: F(R)=1/2 ln (1+R)/(1-R) Then, the rewiring score(rewireTF-gene) was defined, in which values ranges from 0 to 1 and larger value indicates more rewiring effect between TF and gene. rewire_(TF-gene)=P(|X|≤│(F(R_high )-F(R_low ))/√(1.06/n_(high-3) +1.06/n_(low-3) )│ ),X~N(0,1) nhigh and nlow are the sample number in H-group and L-group, separately. Finally, we shuffled the samples’ labels and recalculated the rewire score 1,000 times. We defined the P-value as the fraction of rewire score in shuffled conditions that was larger than in the real conditions. TF-gene dysregulations with BH adjusted P-value < 0.05 were regarded as DMTDs.
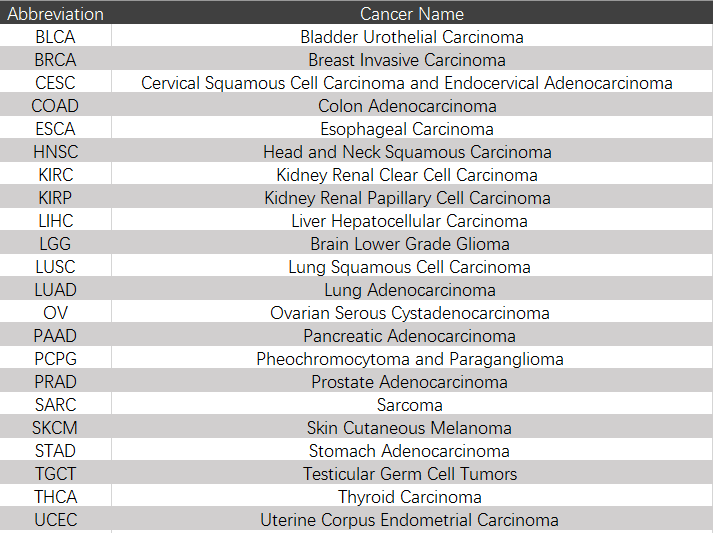
The DMTDB provides an interface to retrieve the information about the DMTDs in cancer. The current release of DMTDB mainly contains the results in 22 kinds of cancer. Users can search by individual TF or by individual gene to view the DMTDs in a certain cancer type. Here, we provide an example where users can click on "examples" and "submit", to check out the information about DMTDs.
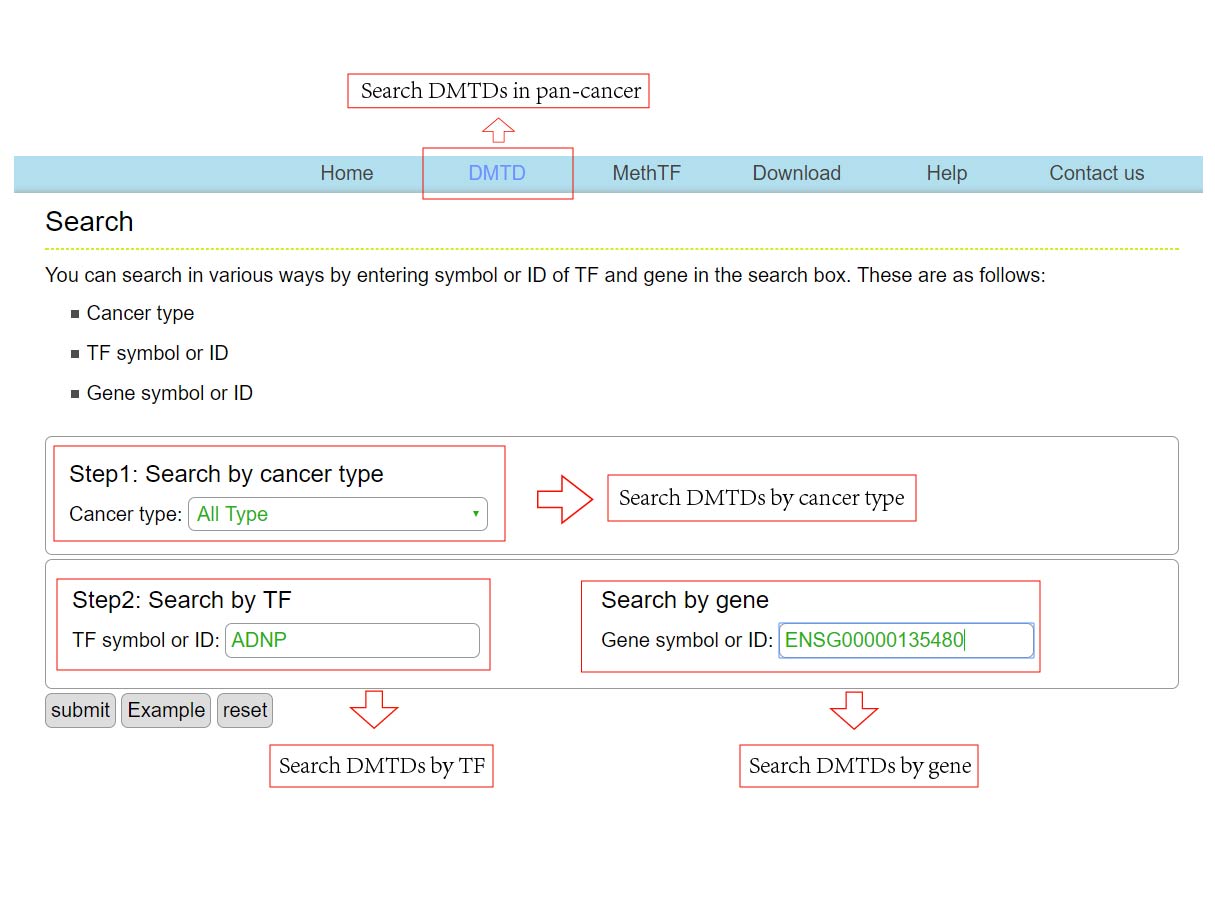
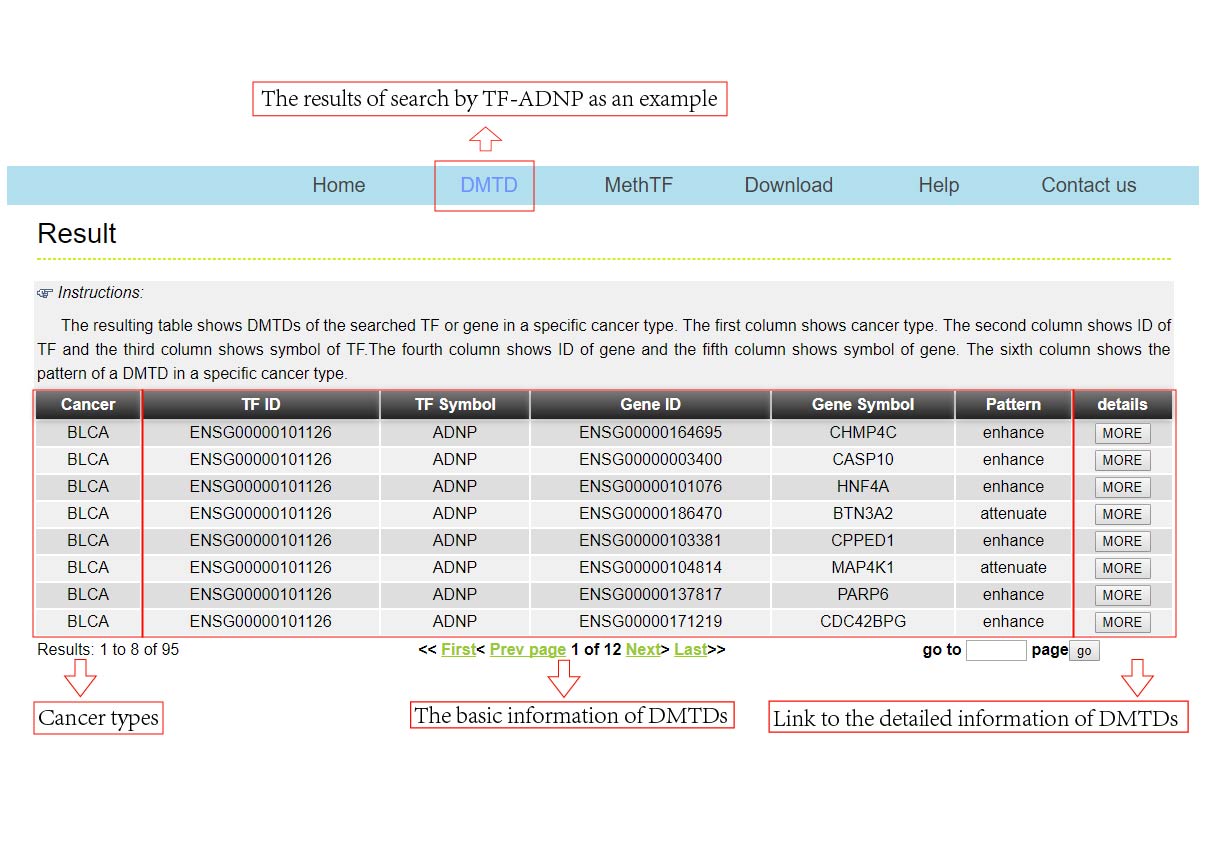
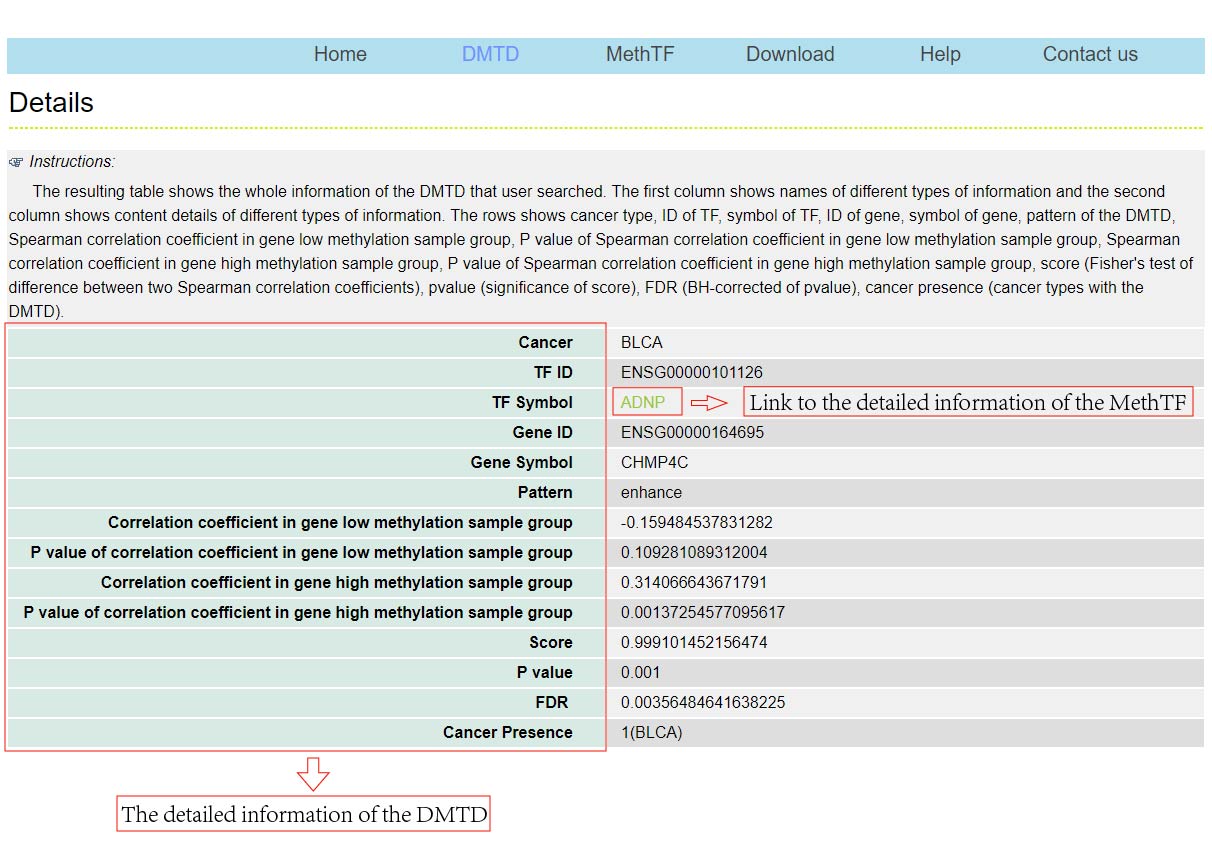
The DMTDB provides an interface to retrieve the information about the MethTFs in cancer. The current release of DMTDB mainly contains the results in 22 kinds of cancer. Users can search by individual TF to view the information of the MethTF in a certain cancer type. Here, we provide an example where users can click on "examples" and "submit", to check out the information about the MethTF.
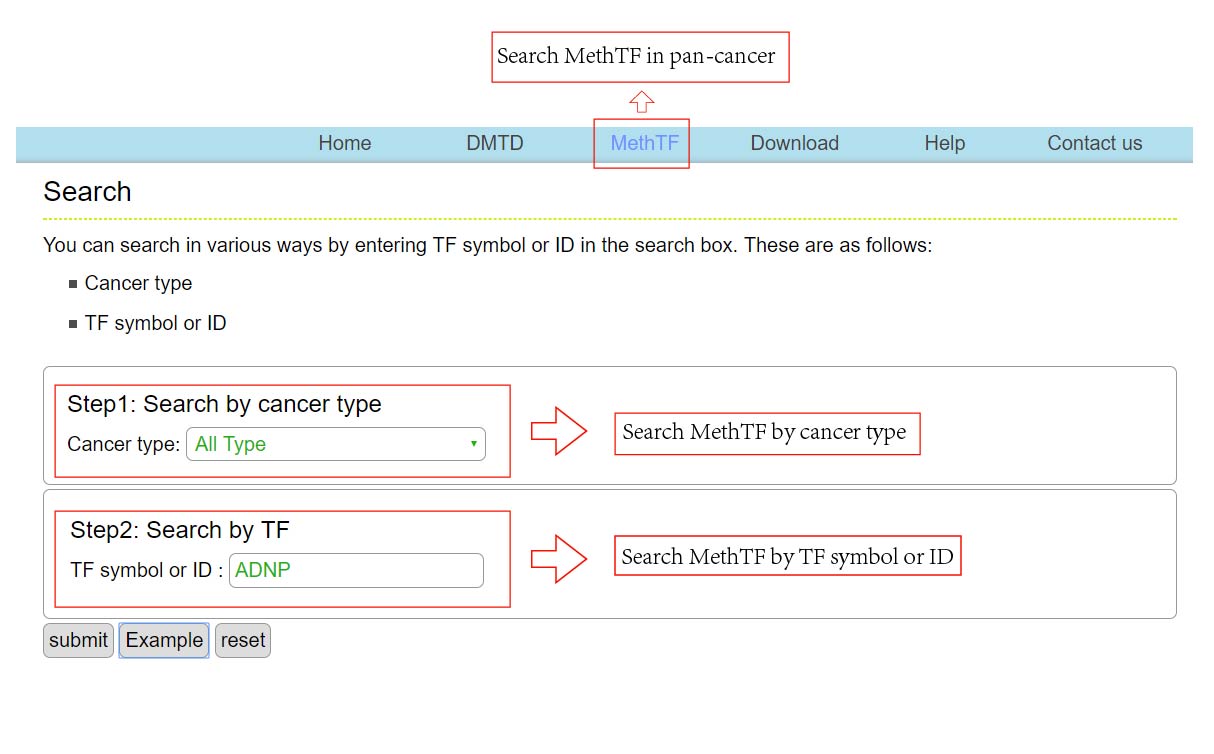
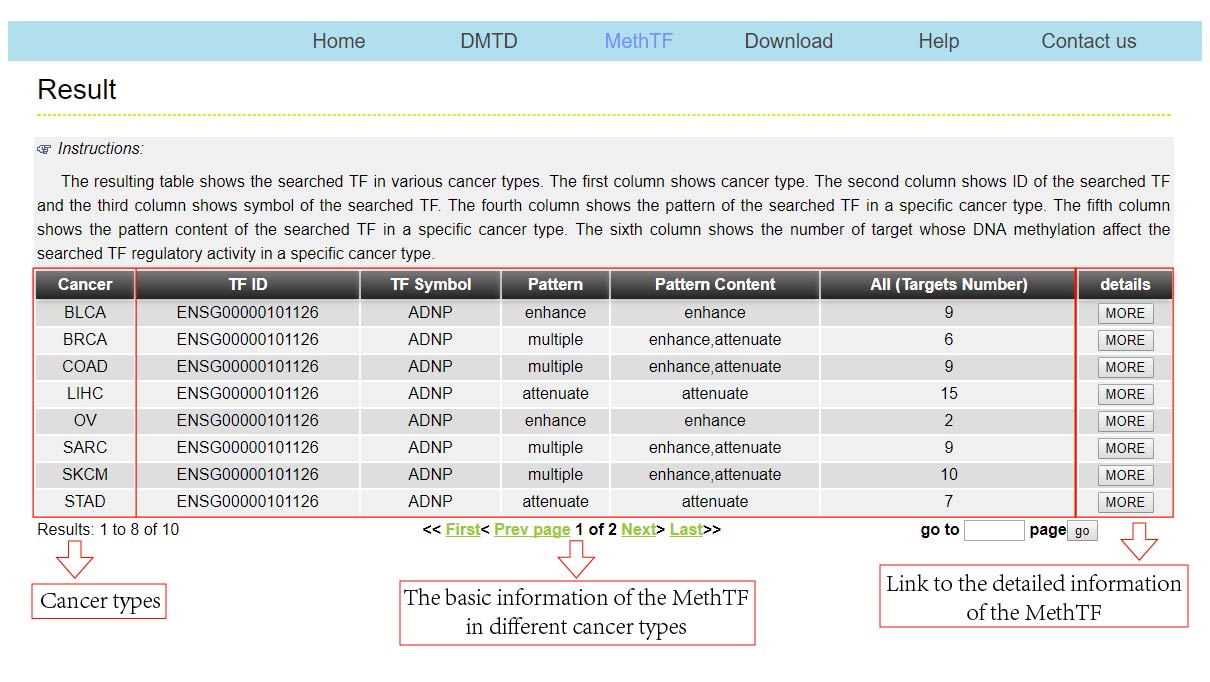
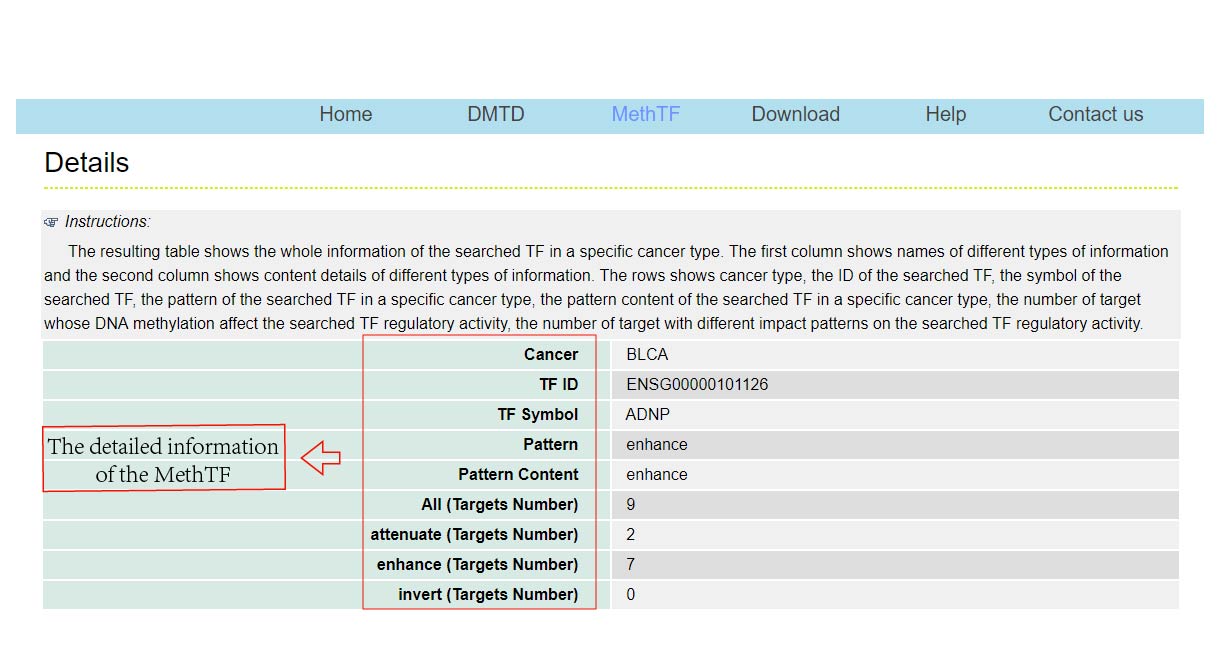
In addition, download links are provided, which allowed you to batch download the information of your interest.
DMTD: DNA methylation - mediated transcription regulation identified based on linear regression model.
MethTF: TFs involved in the DMTD.
DMTD_V2: DNA methylation - mediated transcription regulation are not identified based on linear regression model.
MethTF_V2 : TFs involved in the DMTD_V2.