Database content
MicroEpitope provides a user-friendly searching interface.
Increasing evidences have shown that both bacteria and viruses encode microbial antigens that are molecular mimics of tumour antigens. The burgeoning field of intratumoral microbiome research has been driven by advances in high-throughput detection technologies. Notably, mass spectrometry (MS) technology enables the direct identification of immune epitopes derived from microbiota, extending host immunopeptidome in cancer. Here, we proposed MicroEpitope for providing and visualizing the atlas of HLA-presented immune epitopes derived from cancer microbiome.
1. Human coding protein sequence: we integrated reviewed human protein sequence information from the UniProt (1) database.
2. Human non-coding protein sequence: we referred to IEAtlas (2) database to integrate human non-coding protein sequence information. We integrated Ribo-seq supported ORFs from RPFdb (3), nuORFdb (4) and Translnc (5) with their basic annotations. According to the genome coordinates of ncORFs and the corresponding annotation files, the sequences of ncORFs were obtained by the "getblast" function in R "bedtoolsr" package using the default parameters. All ncORFs with NTG start codons and TAA/TGA/TAG stop codons were kept.
3. For the microbial protein sequence, we collected the literature and microbioTA supporting bacterial species that existed within tumor or in the microenvironment of tumor tissues and integrated the corresponding reviewed protein sequence information of these literature and microbioTA supporting bacterial species from the UniProt database (1,6). We also collected the literature and IEDB support for intratumoral or cancer-associated viral species and integrated the corresponding reviewed protein sequence information of these literature and IEDB supporting viral species from the UniProt database (1,7).
All the available MS-based immunopeptidome datasets were from commonly used proteome databases, including PRIDE (8), MassIVE.quant (9), JPOST (10), iProX (11), PeptideAtlas (12) and Panorama (13). Currently, MicroEpitope reanalyzed 1,190 samples of 24 cancer types.
MicroEpitope further assesses the immunogenicity through several commonly used immunogenic features, including MHC binding affinity, MHC binding stability, and T cell recognition probability. Epitopes with MHC binding affinity ≤ 500 nM, MHC binding stability > 1.4 h, and T cell recognition probability > 1e-16 were defined as immunogenic epitopes (14-16).
Web interface
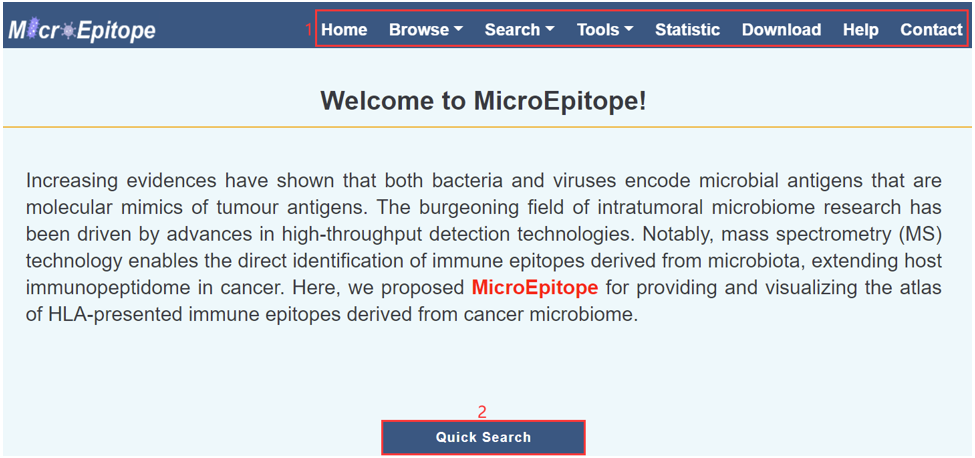
1. Main functions of the database are provided in menu bar form.
2. Click the icon of mouse button to start a quick search.
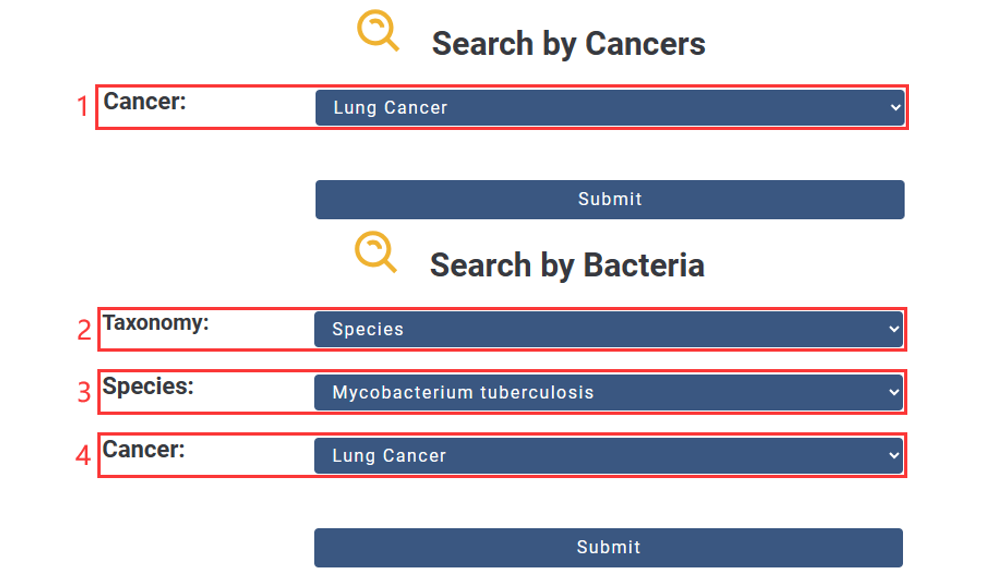
1. Choose the Cancer you are interested in.
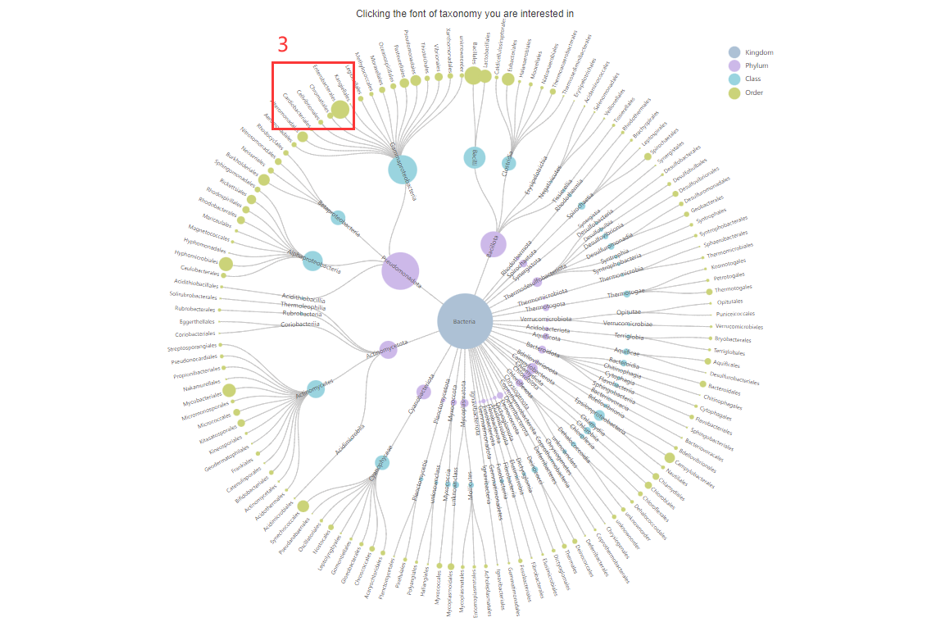
2. Choose the Taxonomy you are interested in.
3. Input Name.
4. Choose the Cancer you are interested in.
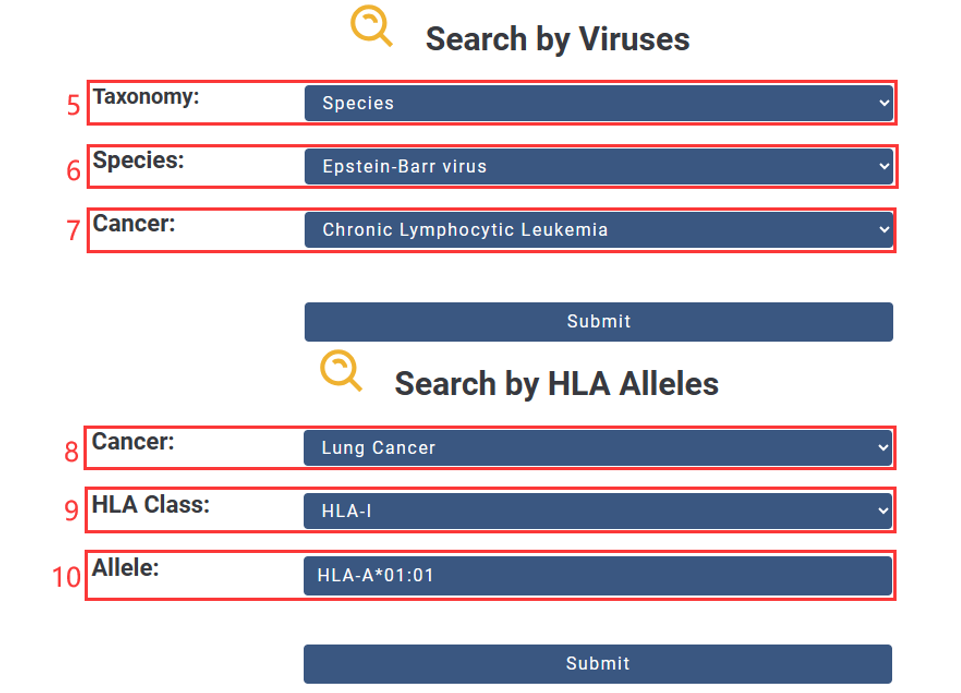
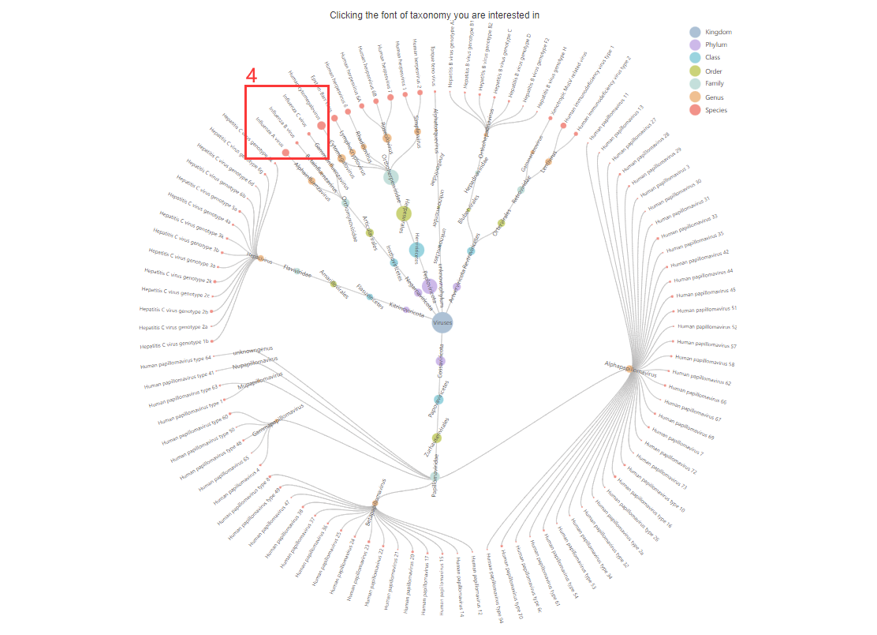
5. Choose the Taxonomy you are interested in.
6. Input Name.
7. Choose the Cancer you are interested in.
8. Choose the Cancer you are interested in.
9. Select HLA Class(HLA-I or HLA-II).
10. Input HLA allele.
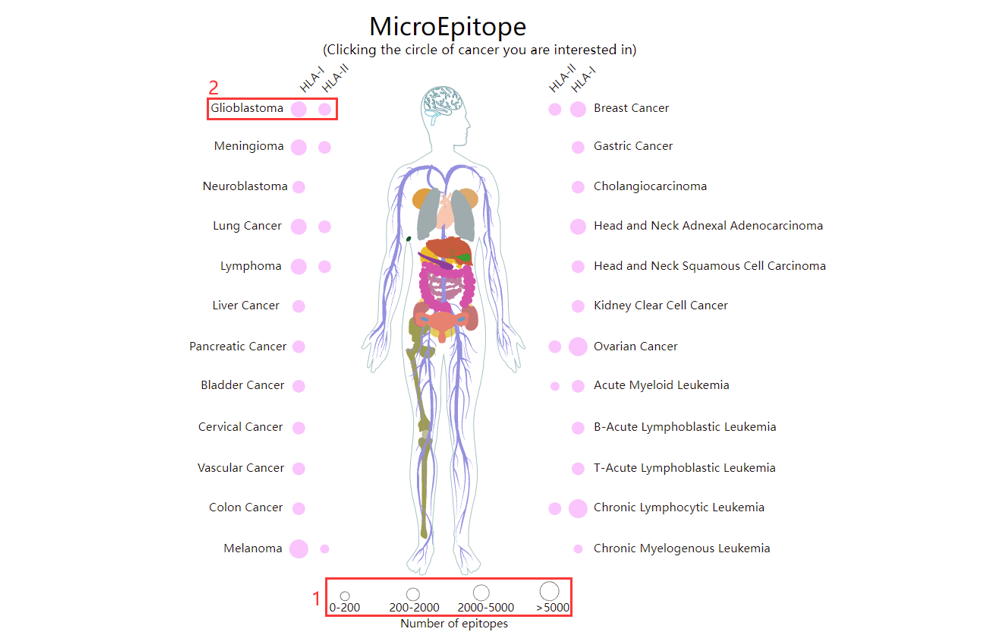
For the browse page, we provide all epitopes indentified by MS-based immunopeptidome across all cancers in MicroEpitope.
1. The size of the dot indicates the number of epitopes in the cancer.
2. Click to browse all epitopes in specific cancer.
3. Click to browse all epitopes in specific taxonomy.
4. Click to browse all epitopes in specific taxonomy.
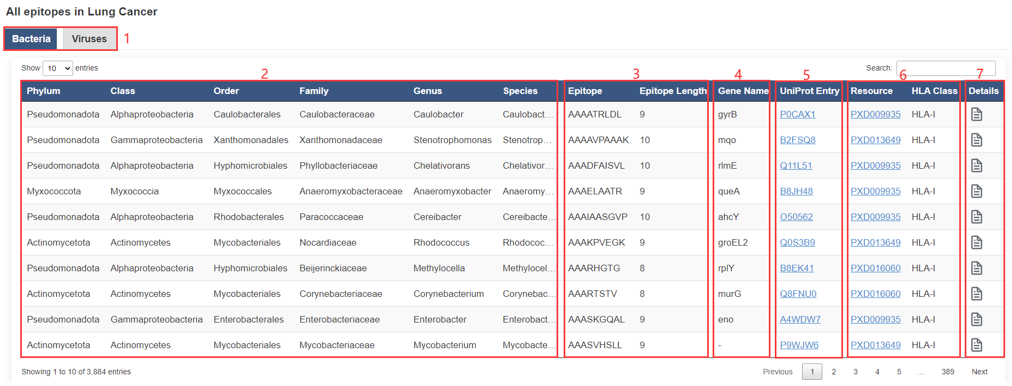
The result page of epitopes is displayed as below.
1. Kingdom.
2. Taxonomy basic information, the "–" symbol indicates that UniProt does not provide or has not annotated the information.
3. Epitope basic informaton.
4. Gene basic information, the "–" symbol indicates that UniProt does not provide or has not annotated the information.
5. Protein basic informaton.
6. Resource.
7. Click to view the detail information.
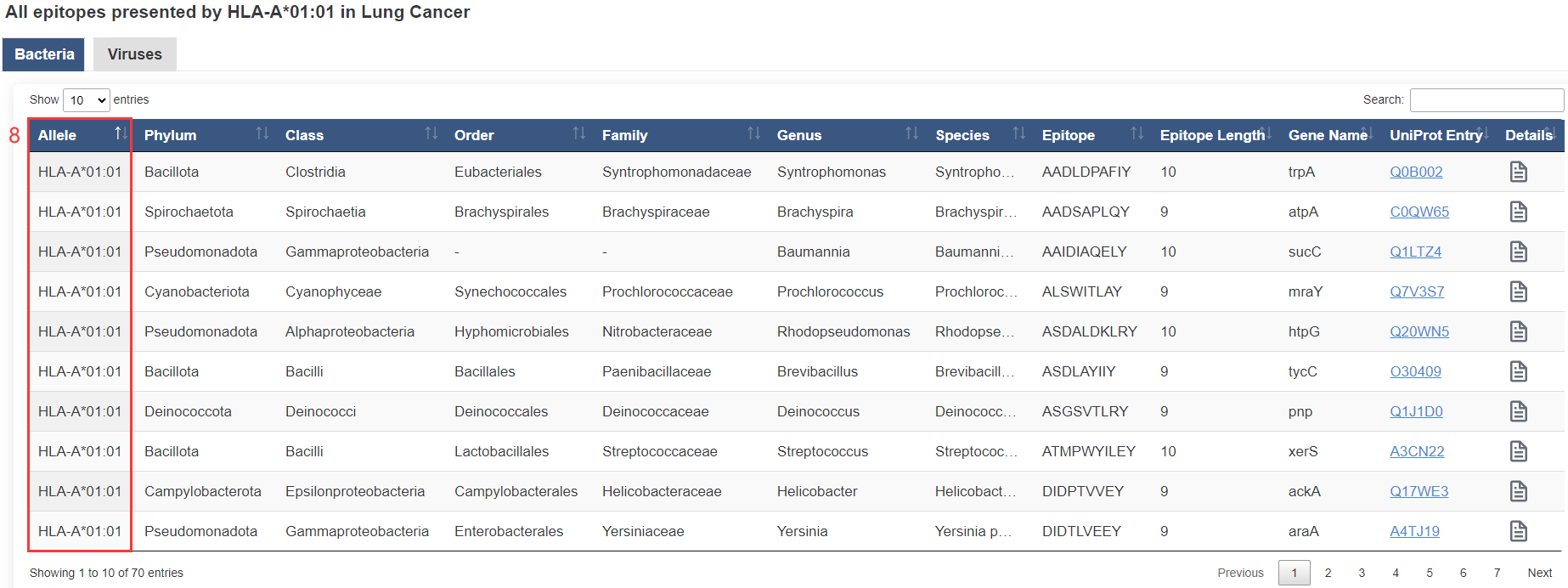
8. HLA allele.
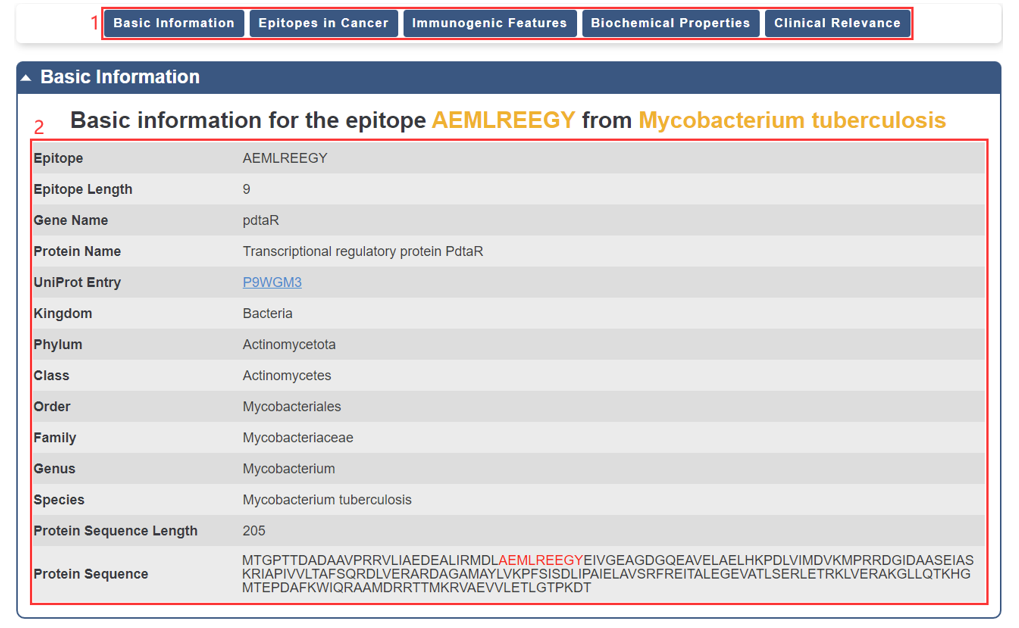
For each epitope, we provide basic information, which is displayed as below.
1. You could directly go to the module of interest by click the axis.
2. The basic information of the epitope.
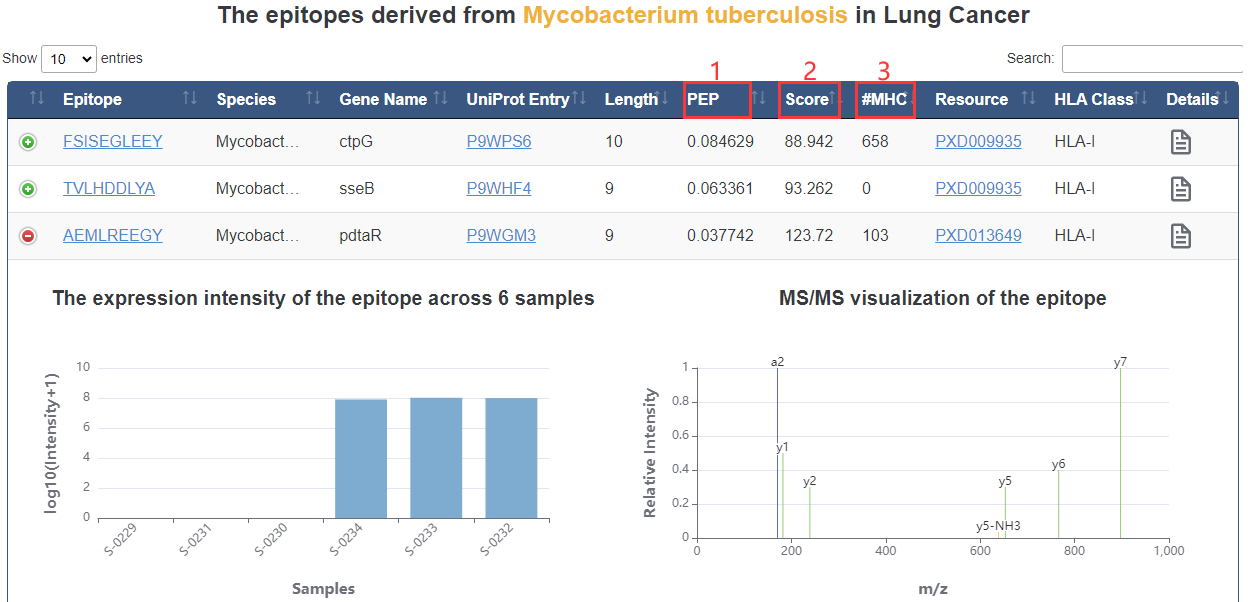
1. PEP: posterior error probability of the identification. This value essentially operates as a p-value, where smaller is more significant.
2. Score: andromeda score for the best associated MS/MS spectrum.
3. #MHC: Number of alleles bound with epitope.
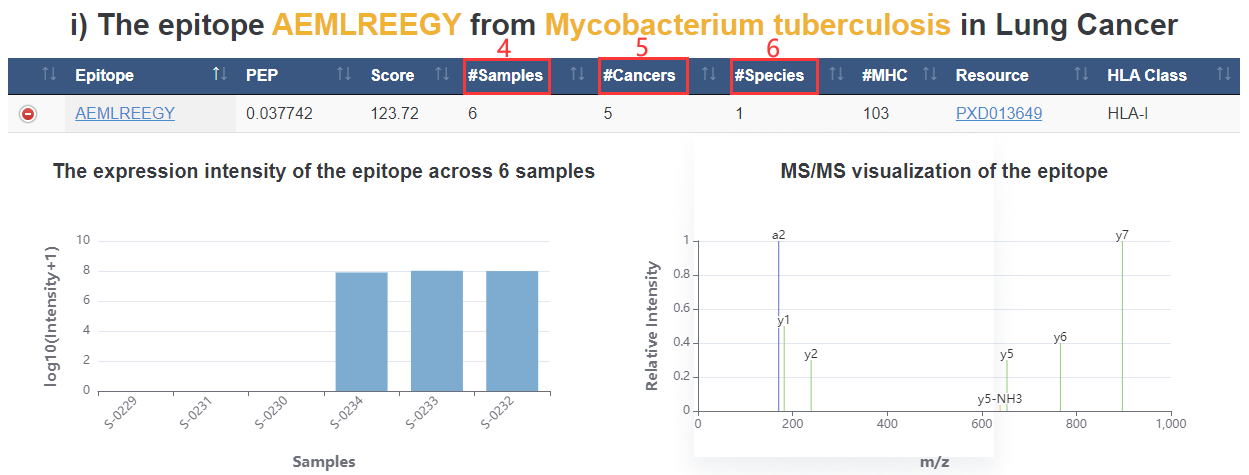
4. #Samples: epitope appears on how many samples.
5. #Cancers: epitope appears on how many cancer types.
6. #Species: epitope appears on how many species types.
7. All epitopes derived from the same species in cancer of interest.
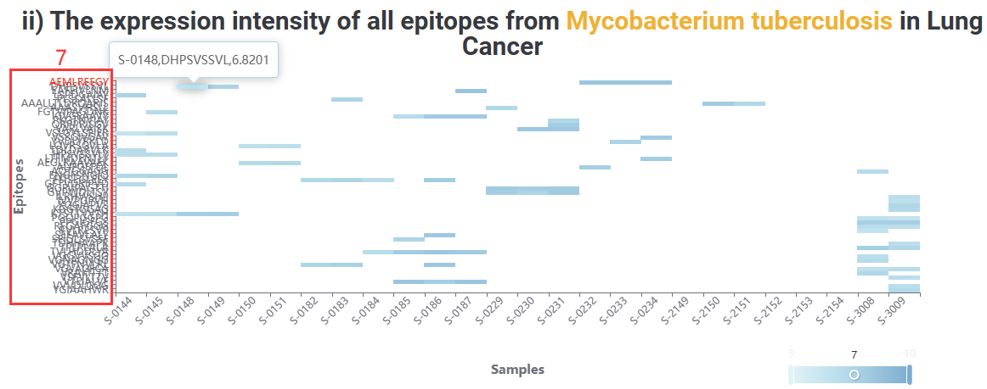
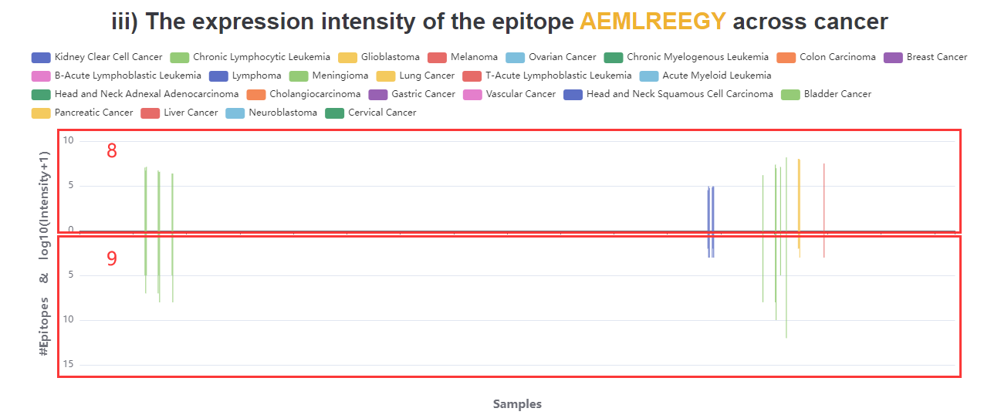
8. The expression intensity of the epitope across all cancers.
9. Number of all epitopes produced by the species producing the epitope in the sample.
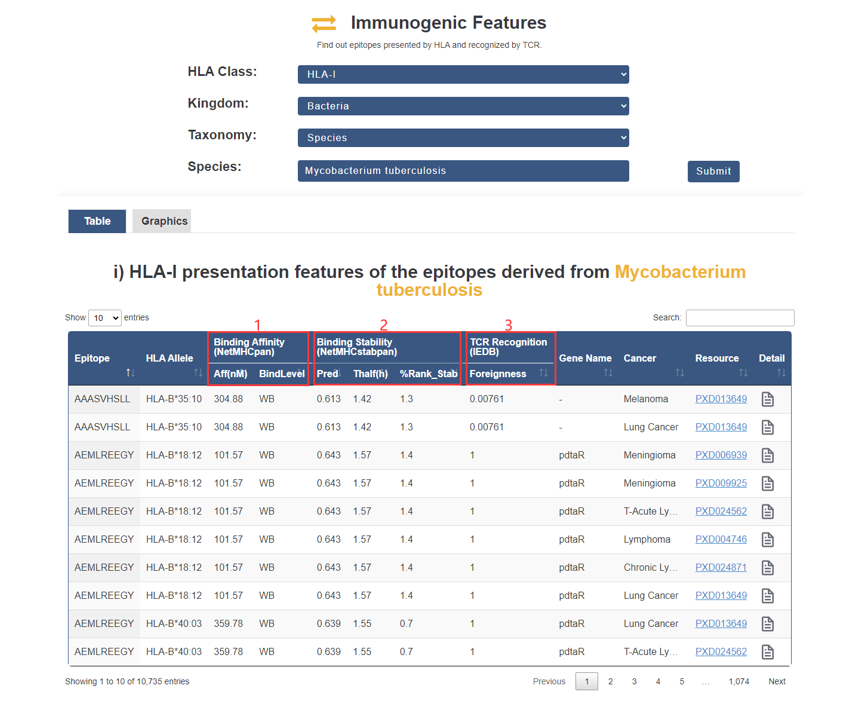
Both presentation features and recognition features of all immunogenic epitopes.
1. Epitope binding affinity was predicted by NetMHCpan and NetMHCIIpan with default settings (either as a strong-binding threshold of 50nM or weak-binding threshold of 500nM).
2. Epitope binding stability was predicted by NetMHCstabpan using default parameters (binding stability greater than 1.4 h).
3. Foreignness: TCR recognition probability derived from homology to known pathogenic peptides in IEDB using the multistate thermodynamic model described by Luksza et al (foreignness more than 1e-16).
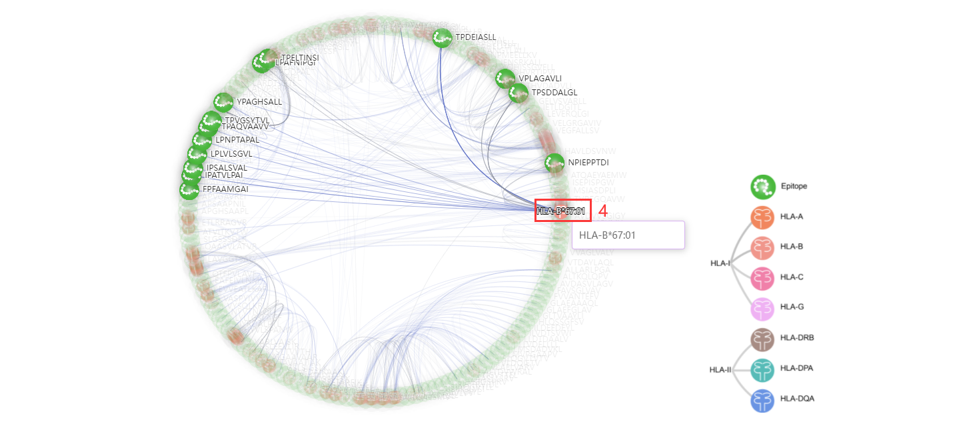
4. Click to enter the AFND database.
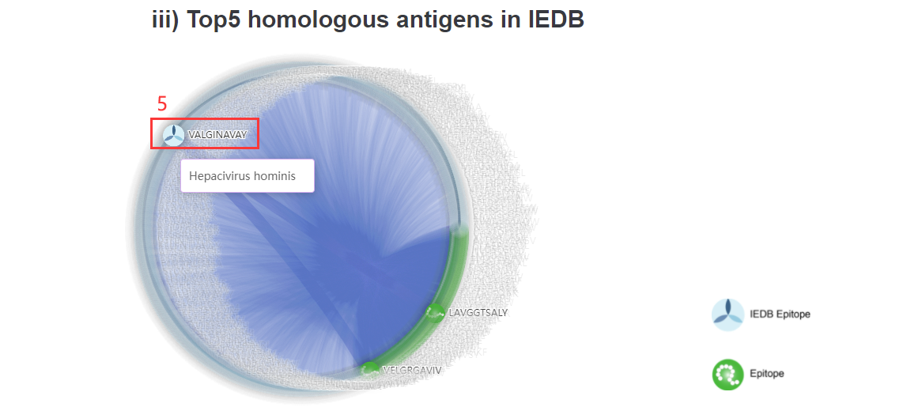
5. Click to the IEDB database. According to the sequence alignment scores, the top5 homologous antigens in IEDB were also shown as a network.
In this page, we provide biochemical properties of all epitopes.
1. Net charge of epitopes when pH=7.
2. Mean hydrophobicity of epitopes.
3. Mean polarity of epitopes.
4. Mean bulkness of epitopes.
5. Boman index of epitopes.
6. Mean aliphatic index of epitopes.
7. Mean pI of epitopes.
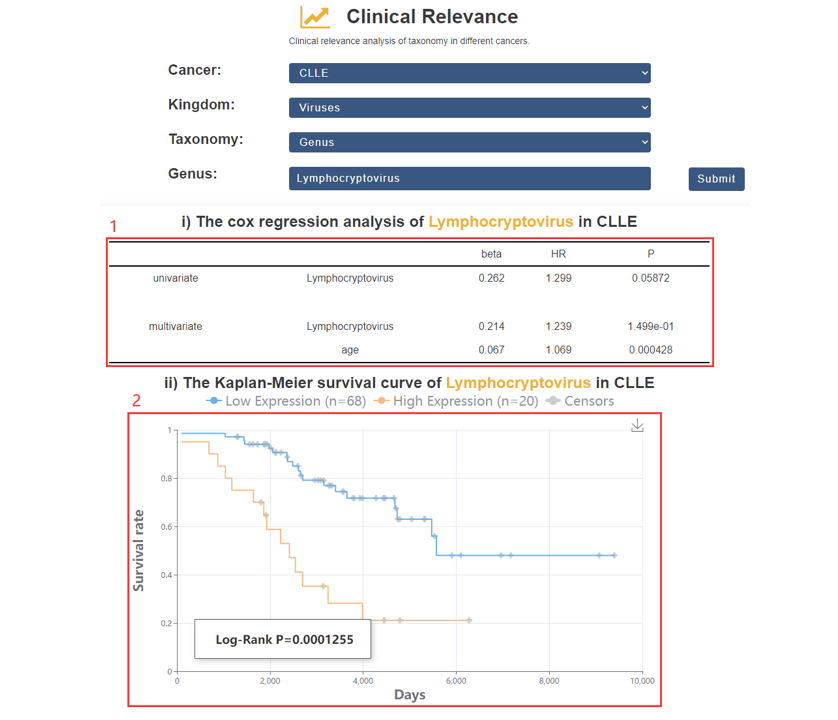
1. The univariate and multivariate Cox regression.
2. Survival analysis.
2. Cai, Y., Lv, D., Li, D., Yin, J., Ma, Y., Luo, Y., Fu, L., Ding, N., Li, Y., Pan, Z. et al. (2023) IEAtlas: an atlas of HLA-presented immune epitopes derived from non-coding regions. Nucleic Acids Res, 51, D409-D417.
3. Wang, H., Yang, L., Wang, Y., Chen, L., Li, H. and Xie, Z. (2019) RPFdb v2.0: an updated database for genome-wide information of translated mRNA generated from ribosome profiling. Nucleic Acids Res, 47, D230-D234.
4. Ouspenskaia, T., Law, T., Clauser, K.R., Klaeger, S., Sarkizova, S., Aguet, F., Li, B., Christian, E., Knisbacher, B.A., Le, P.M. et al. (2022) Unannotated proteins expand the MHC-I-restricted immunopeptidome in cancer. Nat Biotechnol, 40, 209-217.
5. Lv, D., Chang, Z., Cai, Y., Li, J., Wang, L., Jiang, Q., Xu, K., Ding, N., Li, X., Xu, J. et al. (2022) TransLnc: a comprehensive resource for translatable lncRNAs extends immunopeptidome. Nucleic Acids Res, 50, D413-D420.
6. Wang, P., Zhang, S., He, G., Du, M., Qi, C., Liu, R., Zhang, S., Cheng, L., Shi, L. and Zhang, X. (2023) microbioTA: an atlas of the microbiome in multiple disease tissues of Homo sapiens and Mus musculus. Nucleic Acids Res, 51, D1345-D1352.
7. Vita, R., Mahajan, S., Overton, J.A., Dhanda, S.K., Martini, S., Cantrell, J.R., Wheeler, D.K., Sette, A. and Peters, B. (2019) The Immune Epitope Database (IEDB): 2018 update. Nucleic Acids Res, 47, D339-D343.
8. Perez-Riverol, Y., Bai, J., Bandla, C., Garcia-Seisdedos, D., Hewapathirana, S., Kamatchinathan, S., Kundu, D.J., Prakash, A., Frericks-Zipper, A., Eisenacher, M. et al. (2022) The PRIDE database resources in 2022: a hub for mass spectrometry-based proteomics evidences. Nucleic Acids Res, 50, D543-D552.
9. Choi, M., Carver, J., Chiva, C., Tzouros, M., Huang, T., Tsai, T.H., Pullman, B., Bernhardt, O.M., Huttenhain, R., Teo, G.C. et al. (2020) MassIVE.quant: a community resource of quantitative mass spectrometry-based proteomics datasets. Nat Methods, 17, 981-984.
11. Ma, J., Chen, T., Wu, S., Yang, C., Bai, M., Shu, K., Li, K., Zhang, G., Jin, Z., He, F. et al. (2019) iProX: an integrated proteome resource. Nucleic Acids Res, 47, D1211-D1217.
12. Desiere, F., Deutsch, E.W., King, N.L., Nesvizhskii, A.I., Mallick, P., Eng, J., Chen, S., Eddes, J., Loevenich, S.N. and Aebersold, R. (2006) The PeptideAtlas project. Nucleic Acids Res, 34, D655- 658.
13. Sharma, V., Eckels, J., Taylor, G.K., Shulman, N.J., Stergachis, A.B., Joyner, S.A., Yan, P., Whiteaker, J.R., Halusa, G.N., Schilling, B. et al. (2014) Panorama: a targeted proteomics knowledge base. J Proteome Res, 13, 4205-4210.
14. Luo, X., Huang, Y., Li, H., Luo, Y., Zuo, Z., Ren, J. and Xie, Y. (2022) SPENCER: a comprehensive database for small peptides encoded by noncoding RNAs in cancer patients. Nucleic Acids Res, 50, D1373-D1381.
15. Bonsack, M., Hoppe, S., Winter, J., Tichy, D., Zeller, C., Kupper, M.D., Schitter, E.C., Blatnik, R. and Riemer, A.B. (2019) Performance Evaluation of MHC Class-I Binding Prediction Tools Based on an Experimentally Validated MHC-Peptide Binding Data Set. Cancer Immunol Res, 7, 719-736.
16. Wells, D.K., van Buuren, M.M., Dang, K.K., Hubbard-Lucey, V.M., Sheehan, K.C.F., Campbell, K.M., Lamb, A., Ward, J.P., Sidney, J., Blazquez, A.B. et al. (2020) Key Parameters of Tumor Epitope Immunogenicity Revealed Through a Consortium Approach Improve Neoantigen Prediction. Cell, 183, 818-834 e813.