MicroProteinDB provides a user-friendly searching and browsing interface.
A large number of novel ncRNA translation products have been identified and validated. Exploring the structure and functional roles of microprotein encoded by ncRNAs is indispensable. Here, we developed the MicroProteinDB database to provide a knowledge base on sequences, structures and function of ncRNA-derived microprotein.
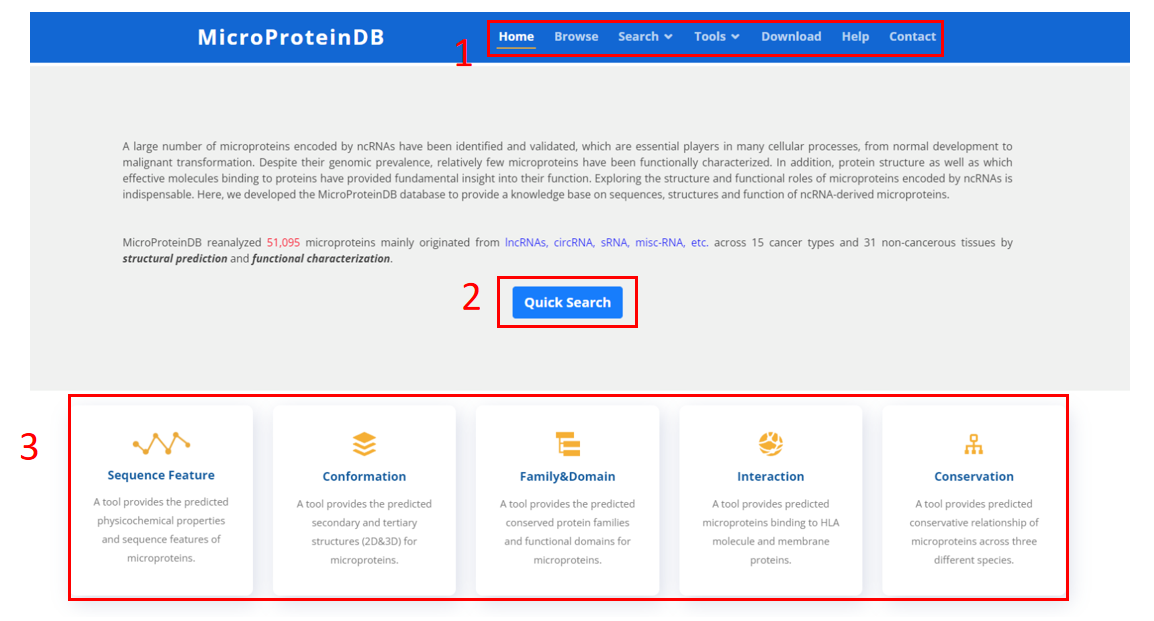
1. Main functions of the database are provided in menu bar form.
2. Click "Start Searching" button to start a quick search.
3. Five kinds of functional tools for MicroProteinDB.
Two ways are provided to query the microprotein encoded by ncRNAs.
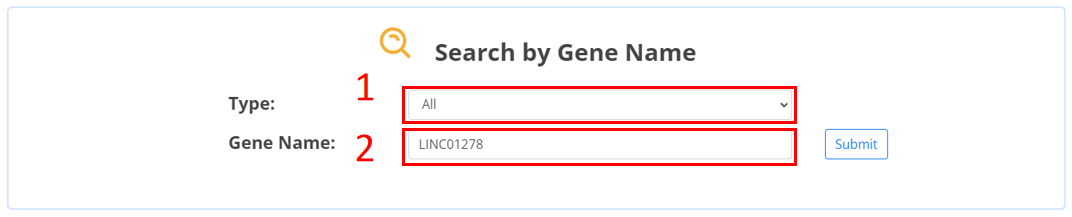
1. Select non-coding RNA types.
2. Input the ncRNA name.
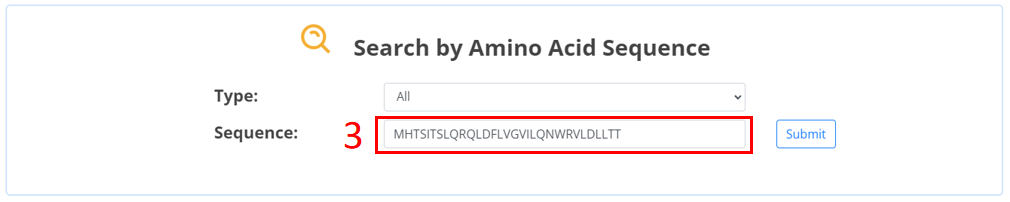
3. Input the amino acid sequence of microprotein.
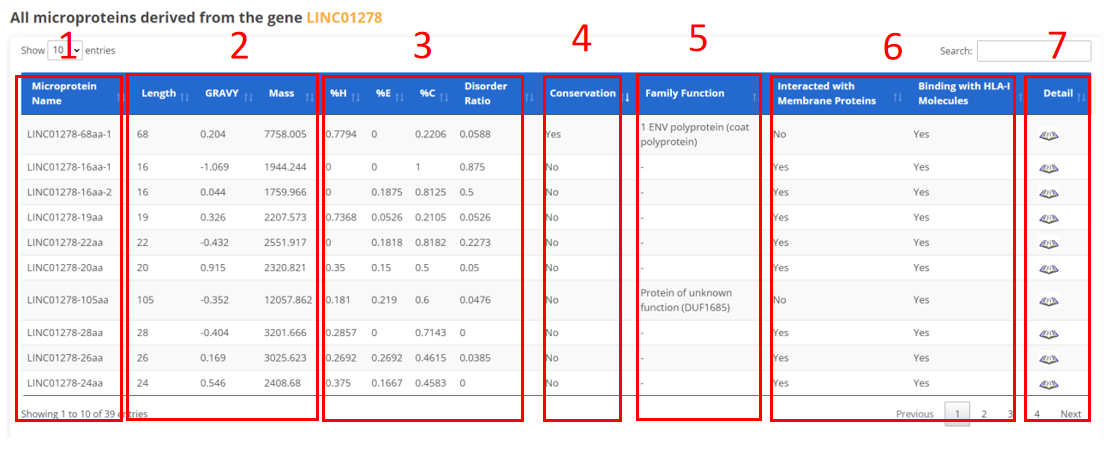
The result page of NcProtein is displayed as below.
1. Microprotein Name.
2. Basic information of microprotein.
3. Microprotein 2D structure and disorder ratio.
4. Conservative relationship of microprotein across three different species.
5. Family and functional domain.
6. Interacting functional molecules.
7. Click to view the detail informations.
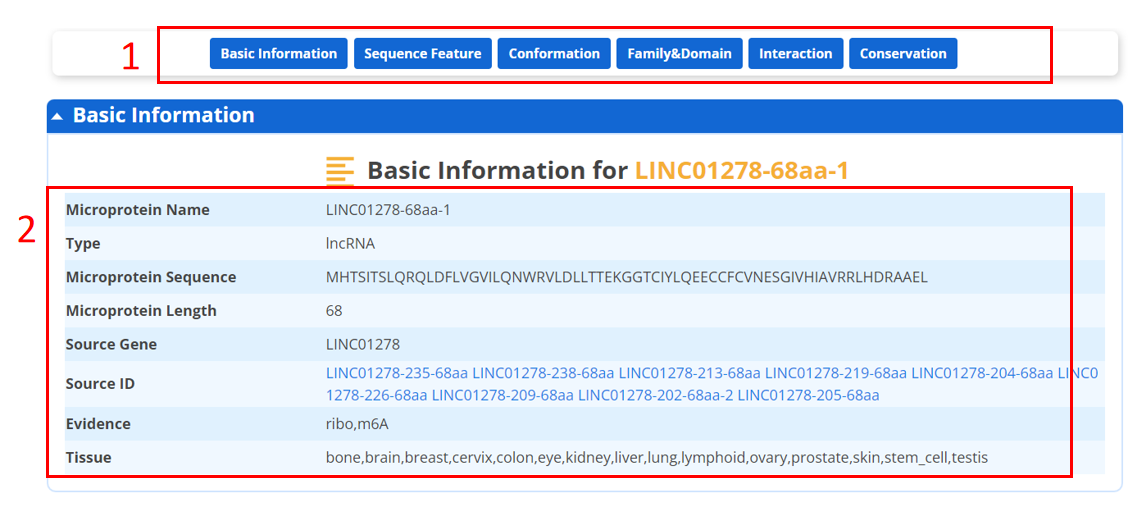
For each microprotein, we provide basic information, which is displayed as below.
1. You could directly go to the module of interest by click the axis.
2. The basic information of the microprotein.
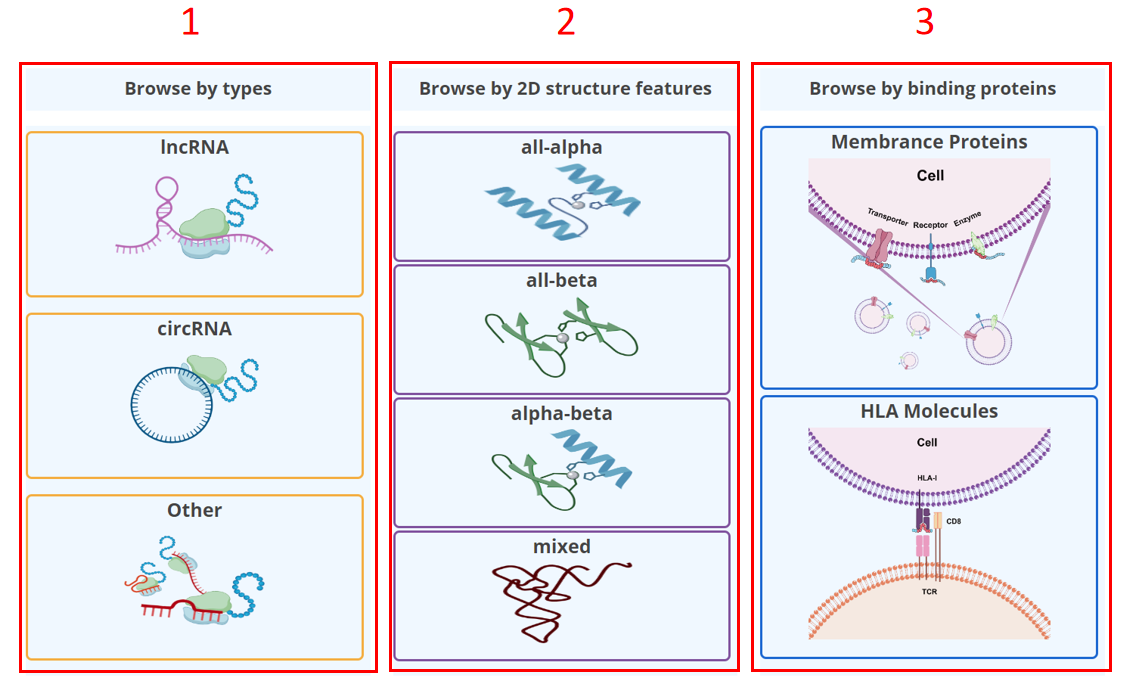
For the browse page, we provide ncRNA encoded microprotein by ncRNA types, 2D structure features, or by peptides bound by MHC I or membrane protein.
1. Browse by ncRNA types.
2. Browse by 2D structure features.
3. Browse by peptides bound by interacting functional molecules.
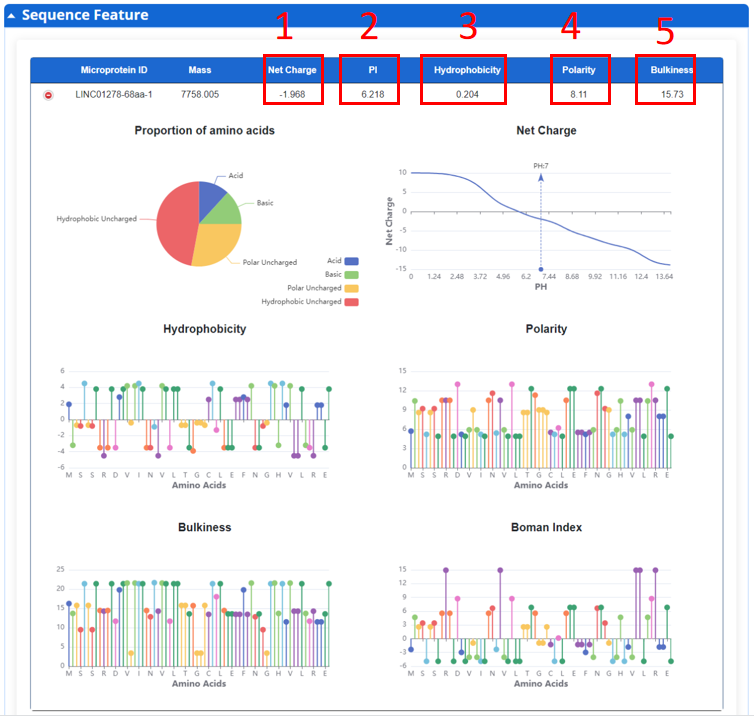
In this page, we provide biochemical properties of all microprotein.
1. Net Charge of microprotein when pH=7.
2. Mean hydrophobicity of microprotein.
3. Mean polarity of microprotein.
4. Mean bulkness of microprotein.
5. Boman Index computes the potential protein interaction index proposed by Boman (2003) based in the amino acid sequence of a protein.
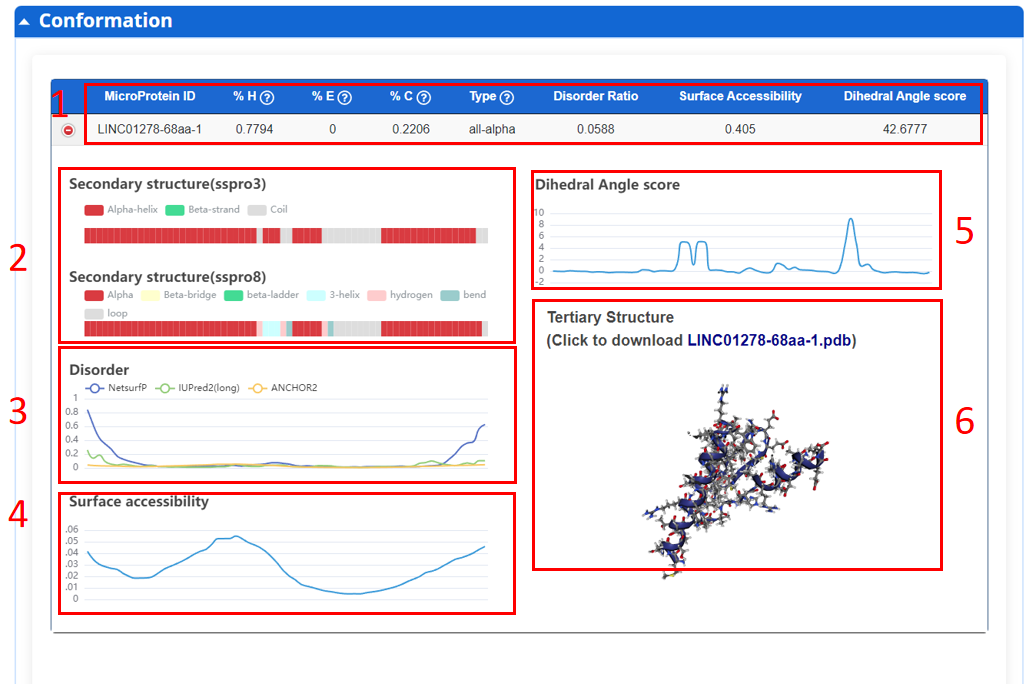
In this page, we provide conformation including 2D and 3D structure of all microprotein.
1. 2D and 3D structure features.
2. Secondary structure.
3. Disorder score.
4. Surface accessibility.
5. Dihedral angle score.
6. Visualization of 3D structure and pdb data download.
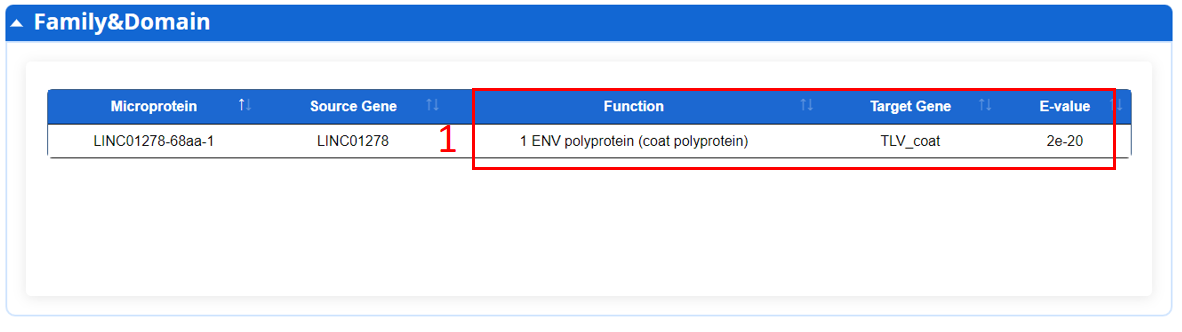
For the microprotein, we provide family and functional domains of microprotein encode by ncRNA according to Pfam database.
1. Function description.
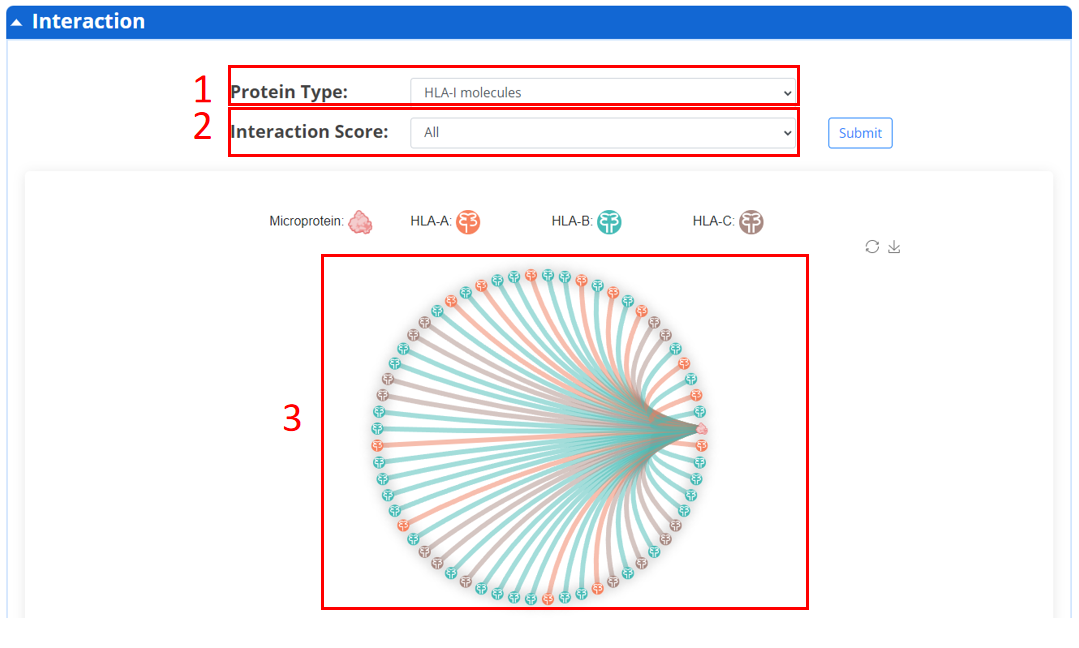
In this page, we provide candidate functional microproteins binding major histocompatibility complex (MHC) class I proteins and five classes of membrane proteins.
1. Types of interacting proteins.
2. Interaction score.
3. Graphic visualization results of interaction results.
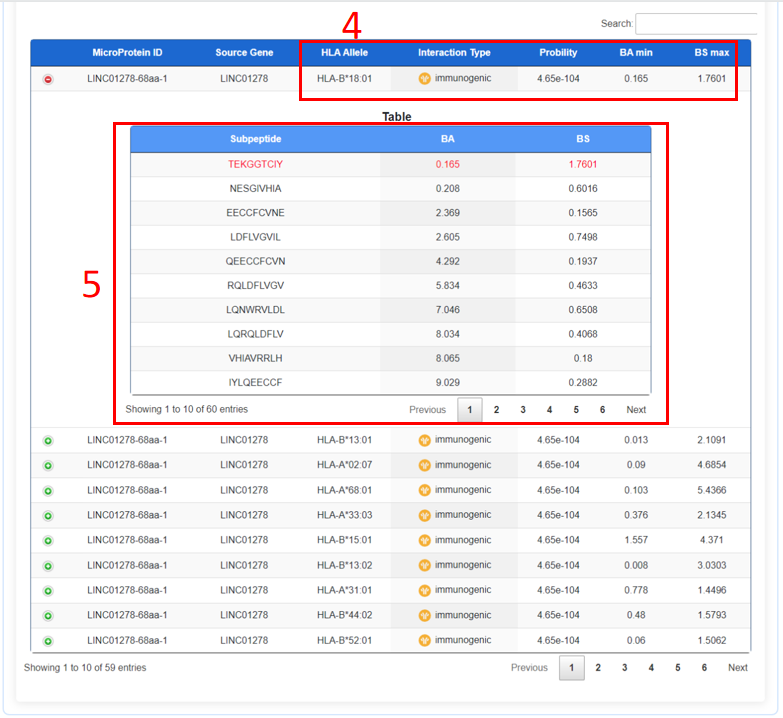
4. Binding HLA molecules and corresponding scores.
5. The minimal 9 amino acid binding core directly in contact with the MHC I.
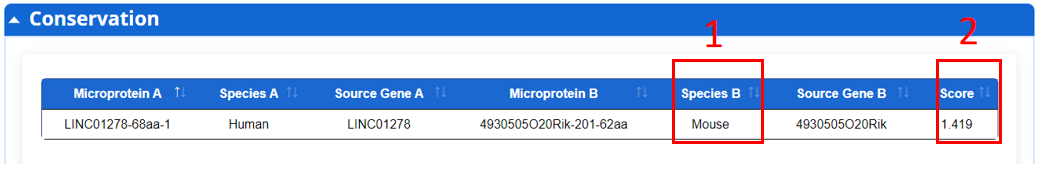
For the microprotein, we provide conservative relationship of microprotein encode by ncRNA across three different species.
1. Species.
2. Conservation score.