1.Overview of PRES
1.1 Overview of PRES
PRES, a webserver decode the functional perturbations of RNA Editing Sites based on Editome profiling.
in the downstream functional interpretation of RNA Editome, with no need for programming skills.
After users uploading an editome profile in samples of different groups,
PRES will first finish annotation of various genomic elements and detect differential-editing sites under the user-selected method and threshold.
Next, the downstream functional perturbations of differential-editing sites will be interpreted from four different levels:
i) gain or loss regulations of miRNAs or RNA-binding proteins;
ii) RNA secondary structure changes or locating at the protein structural region;
iii) changes of amino acid composition by non-synonymous editing sites;
v) the significantly enriched biological functions or pathways.
Finally, genes with the top-scoring of functional perturbation degree will be listed to assist the user for further investigation.
PRES provides both tables and visualization analytical results, and three tools are also respectively developed for independent function analysis.
1.2 The list of databases and resources integrated in PRES
| Name | Web Site | miRNA | RBP | Gene | RNA Secondary Structure | domain | disorder |
|---|---|---|---|---|---|---|---|
| ANNOVAR | https://annovar.openbioinformatics.org/en/latest/ | √ | |||||
| miRBase | https://www.mirbase.org/ | √ | |||||
| miRanda | http://www.microrna.org/microrna/home.domiRanda | √ | |||||
| TargetScan | https://www.targetscan.org/vert_80/ | √ | |||||
| MEME | https://meme-suite.org/meme/ | 160 RBP 3526 motif | |||||
| UCSC | http://genome.ucsc.edu/ | gene sequence | |||||
| GenCode h19 | https://www.gencodegenes.org/ | gene annotation | |||||
| ANCHOR2 | http://anchor.elte.hu/ | √ | |||||
| pfamscan | https://www.ebi.ac.uk/Tools/pfa/pfamscan/ | √ | |||||
| RNAfold | http://rna.tbi.univie.ac.at/cgi-bin/RNAWebSuite/RNAfold.cgi | √ | |||||
| ATtRACT | https://attract.cnic.es/ | √ |
2.How to use PRES?
2.1 Input
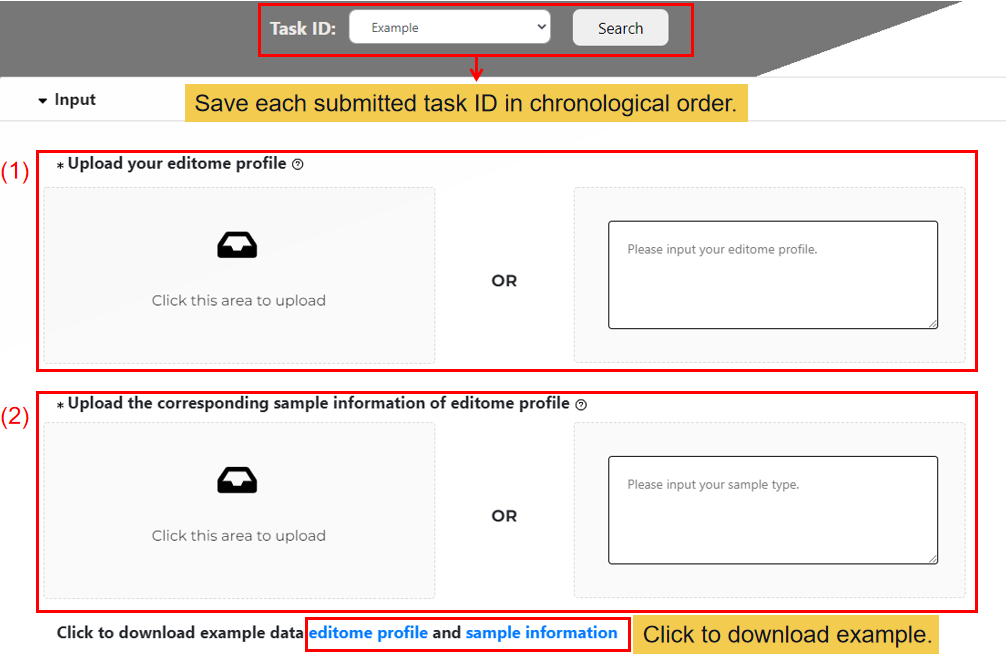
(1) Upload your editome profile.
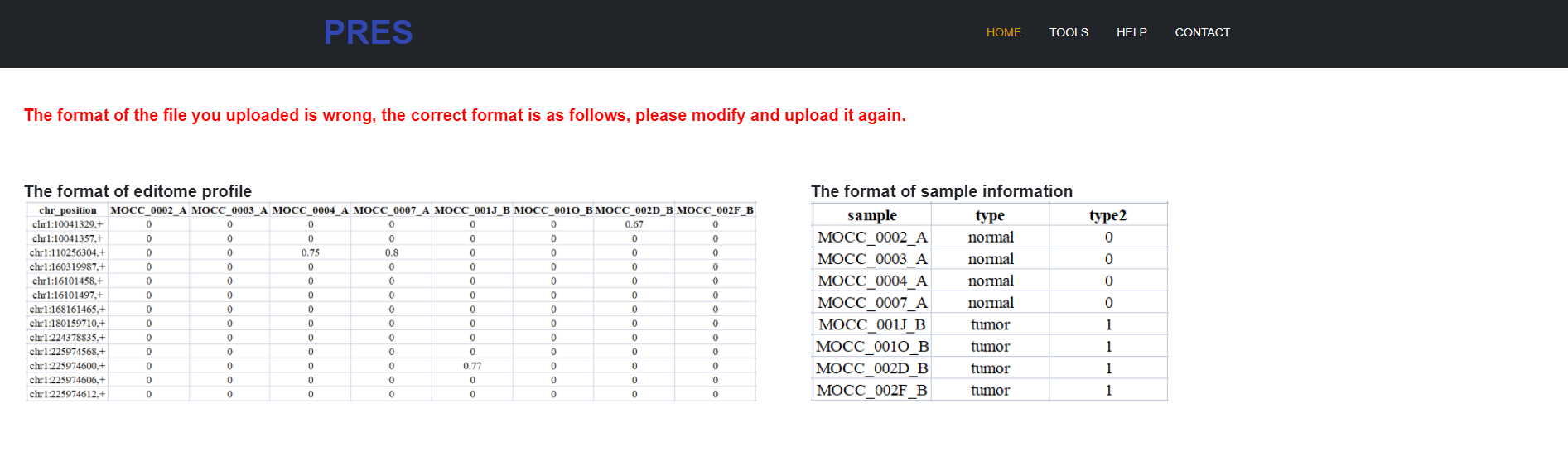
Firstly, uploading your editome profile: Editing profile: column 1 = editing sites (for example: chr1:100127891,+); row 1 = samples labels; no missing values.
(2) Upload the corresponding sample information of editome profile.
And then uploading your sample type data or enter a list of related sample types (either the uploaded file or the input list).
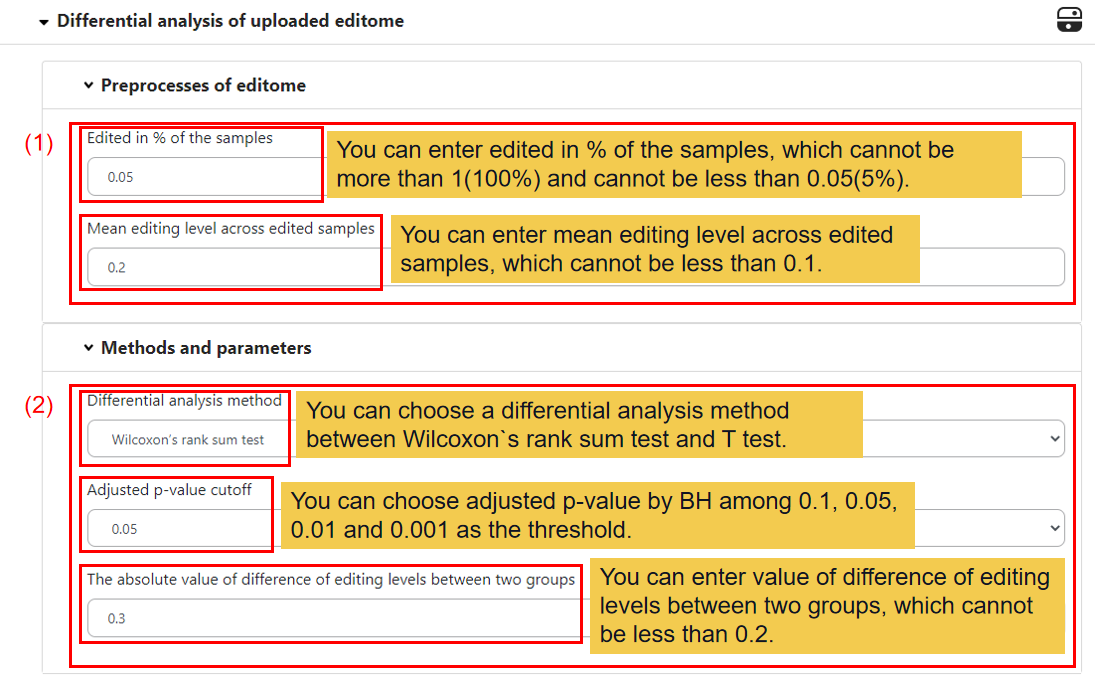
2.2 Differential analysis of uploaded editome
In this module, you can set your own filtering threshold.
(1) Preprocesses of editome.
You can set the min sample numbers the editing site edited (e.g. 0.05) and the min mean editing levels of the editing site across edited samples (e.g. 0.2).
(2) Methods and parameters.
You can set the differential analysis method between the two groups, default is Wilcoxon’s rank sum test, you can also choose T test, and the adjust p-value cutoff of the test (e.g. 0.05). And then you can set the absolute value of difference of editing levels between two groups (e.g. 0.3).
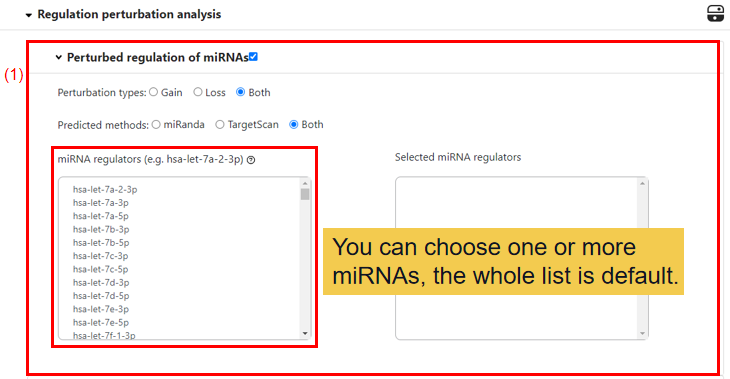
2.3 Regulation perturbation analysis.
(1) Perturbed regulation of miRNAs.
You can choose the perturbation type of miRNA, default is Both, allowed values include Gain and Loss. Then the method to predicted the perturbation type also can be chosen, default is Both, allowed values include Mirnada and TargetScan. Lastly, you can select one miRNA or more miRNAs or all for following analysis.
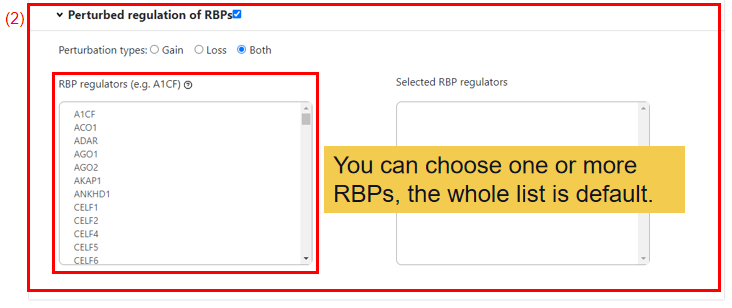
(2) Perturbed regulation of RBPs.
You can choose the perturbation type of RBPs, default is Both, allowed values include Gain and Loss. And then you can select one RBP or more RBPs or all for following analysis.
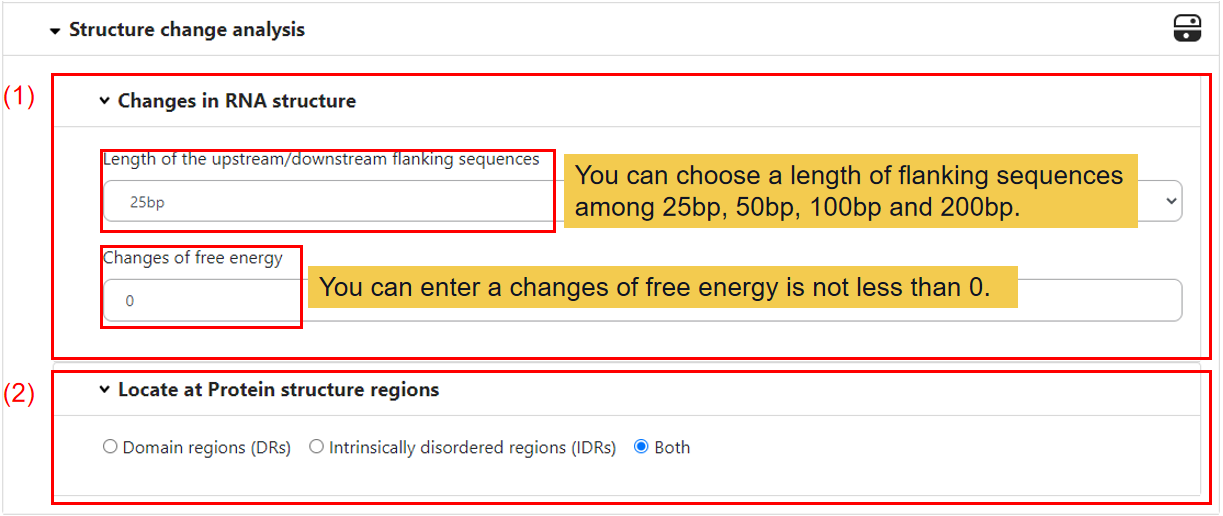
2.4 Structure change analysis.
(1) Changes in RNA structure.
Firstly, you can select the length of the upstream/downstream flanking sequences, default is 25bp, and the changes of free energy after edited, default is 0.
(2) Locate at protein structure regions.
If the editing site locates in the exonic region of mRNA, the amino acids sequence may be changed. And we have predicted where the editing amino acid located at protein structure regions, default is Both, allowed values include Domain regions (DRs) and Intrinsically disordered regions (IDRs).
2.5 Run!
Upload and run!

3.What will you get from PRES?
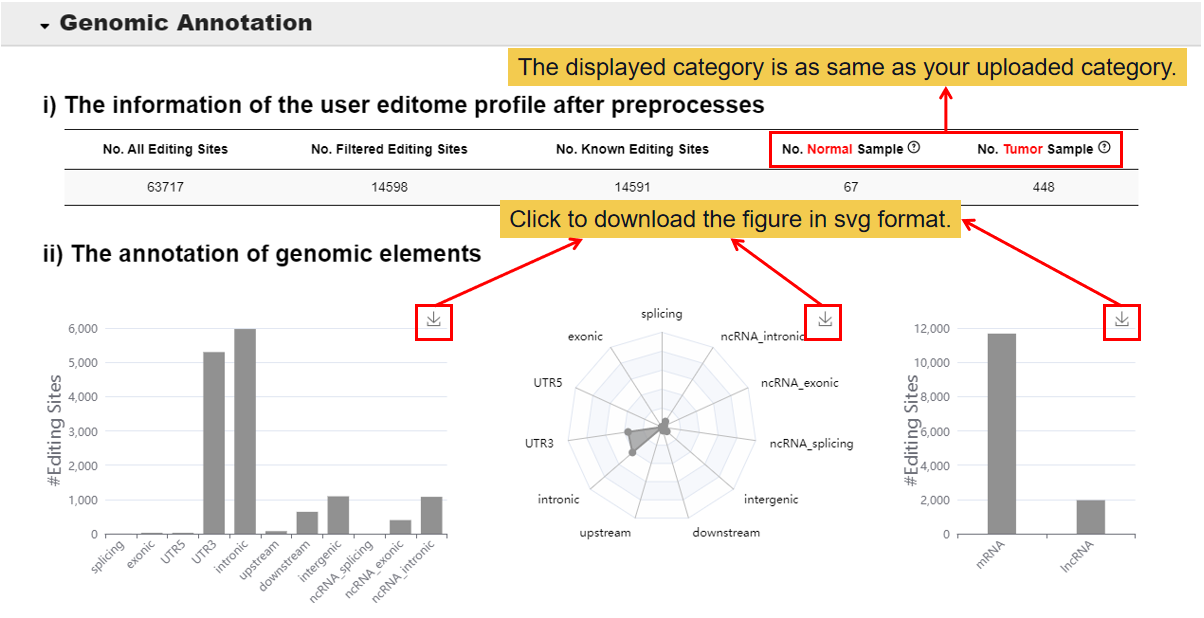
3.1 Genomic Annotation.
In this module, you will get the information of the user editome profile after preprosses and the annotation of genomic elements.
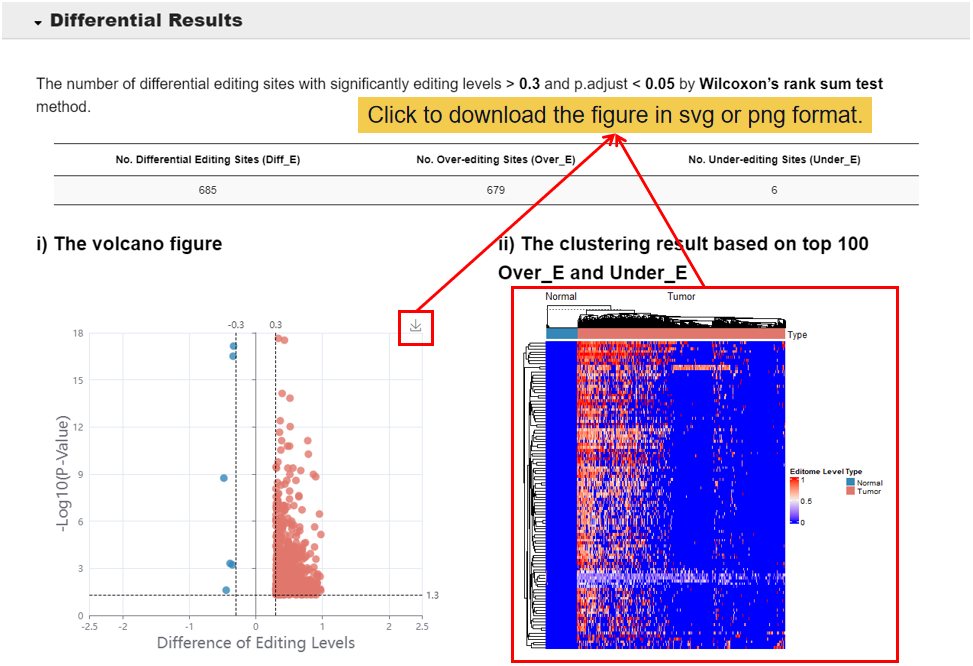
3.2 Differential Results.
This module shows the differential results of the user editome profile and visualizing the differential editing site, including 1) the volcano figure of the differential editing site, 2) the clustering result based on top 100 over-editing sites and under-editing sites, 3) the number and enrichment of differential editing site, 4) the annotation and enrichment results of Diff_E in genomic elements, 5) the table result of differential editing sites.
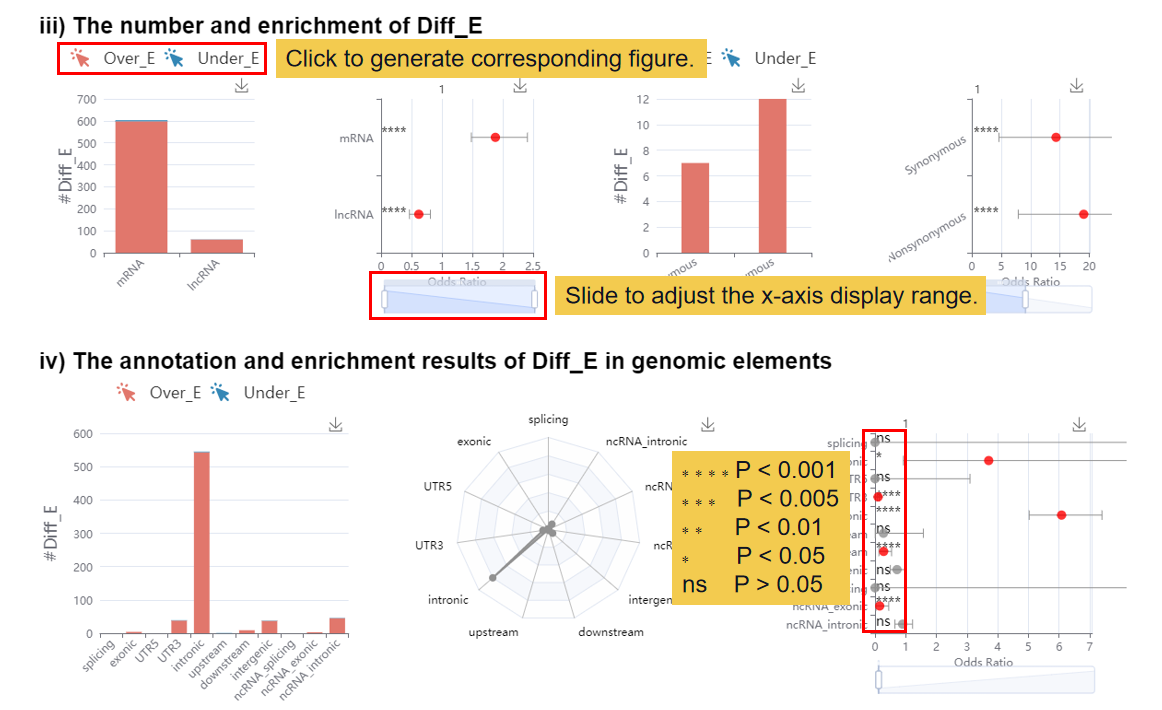
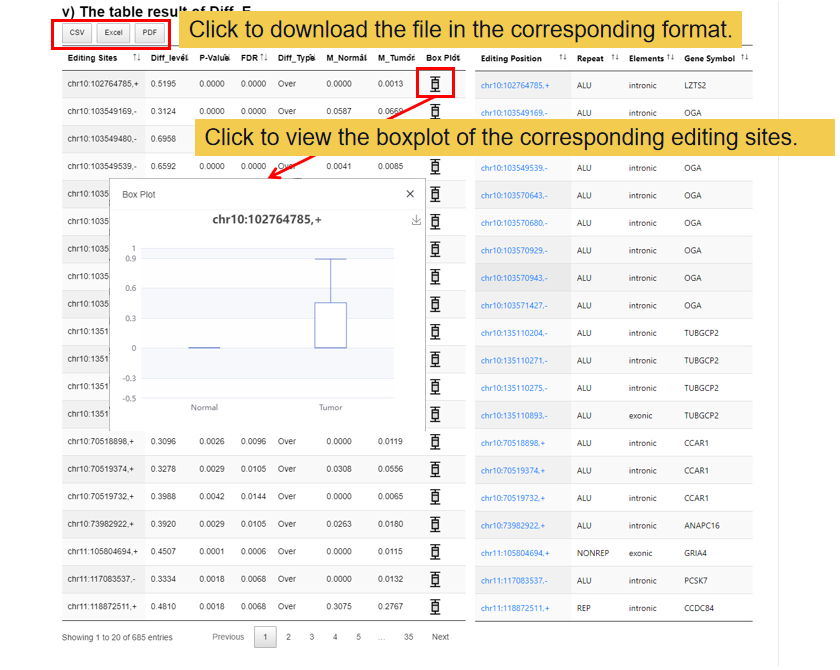
3.3 Functional Perturbations.
3.3.1 Perturbed regulation
In this module, you can get two functional perturbations of RNA editing sites, miRNA and RNA binding protein (RBP). At present, two methods we have used to predict the functional perturbations of RNA editing sites with miRNAs, miRnada and TargetScan, and one for RBPs, MEME. Gain: there is no functional perturbation before edited but gain after edited; Loss: there are some functional perturbations of RNA editing sites with miRNAs/RBPs before edited but loss after edited.
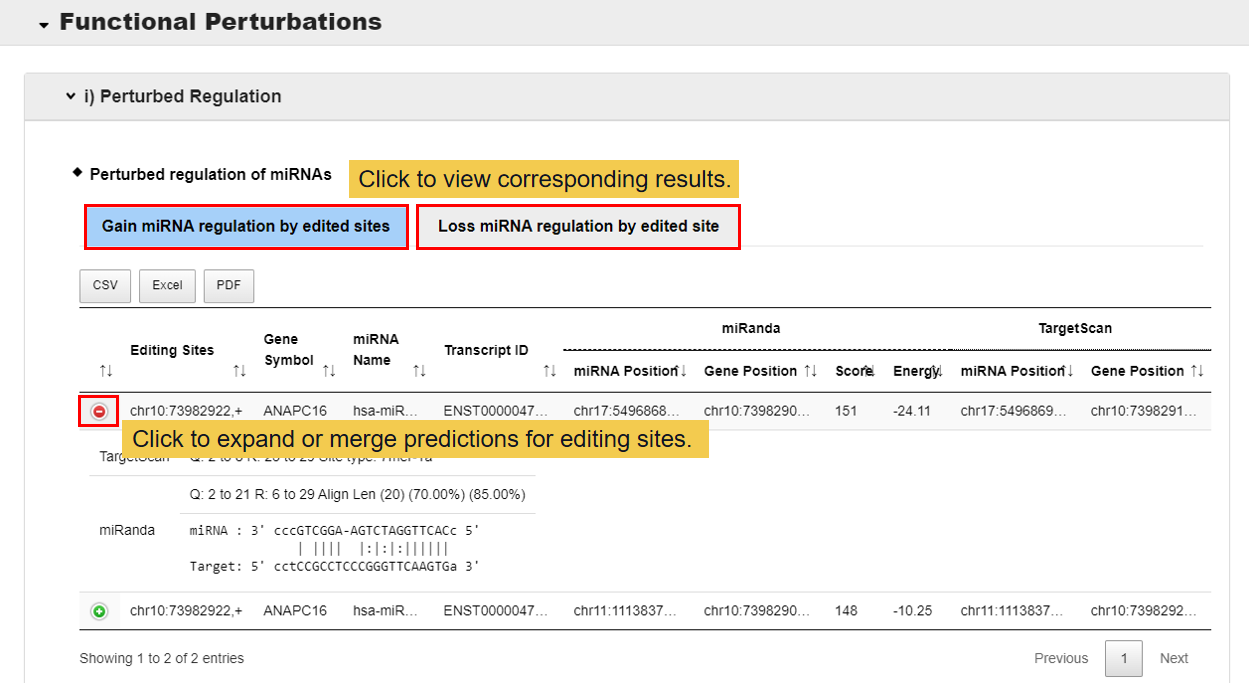
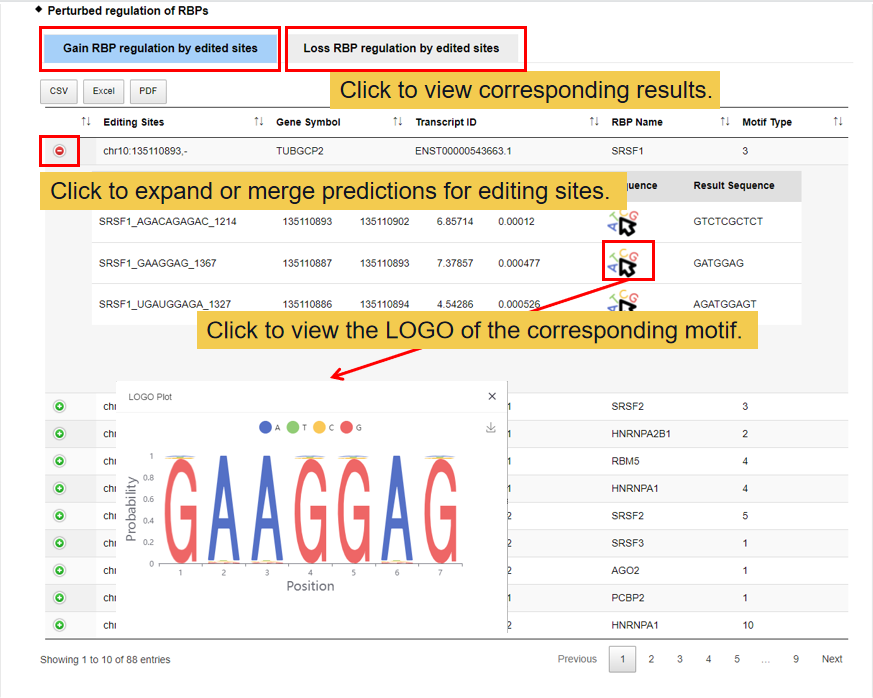
3.3.2 Structure change analysis.
This module shows whether the RNA structure changes after edited, and where the editing RNA sites located at protein structure regions.
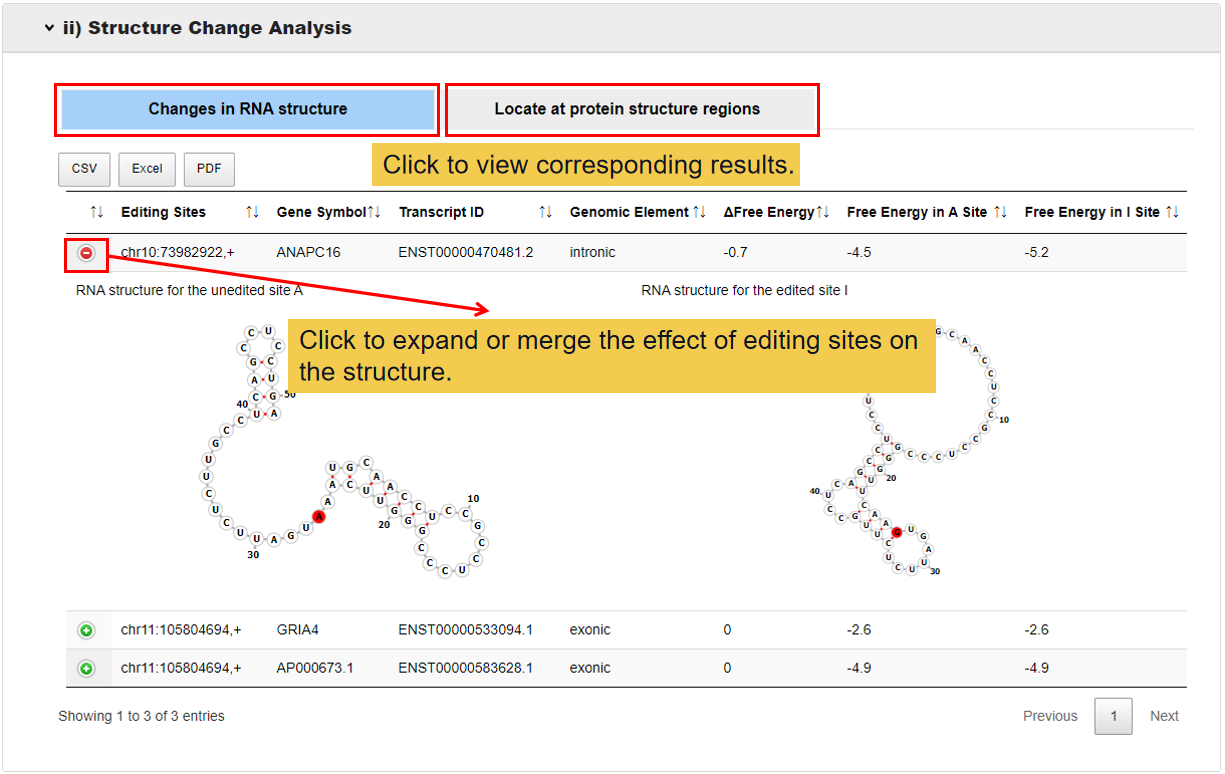
3.3.3 Changes of amino acid composition.
In this module,whether the amino acids changes after edited and where the editing amino acid located at protein structure regions.
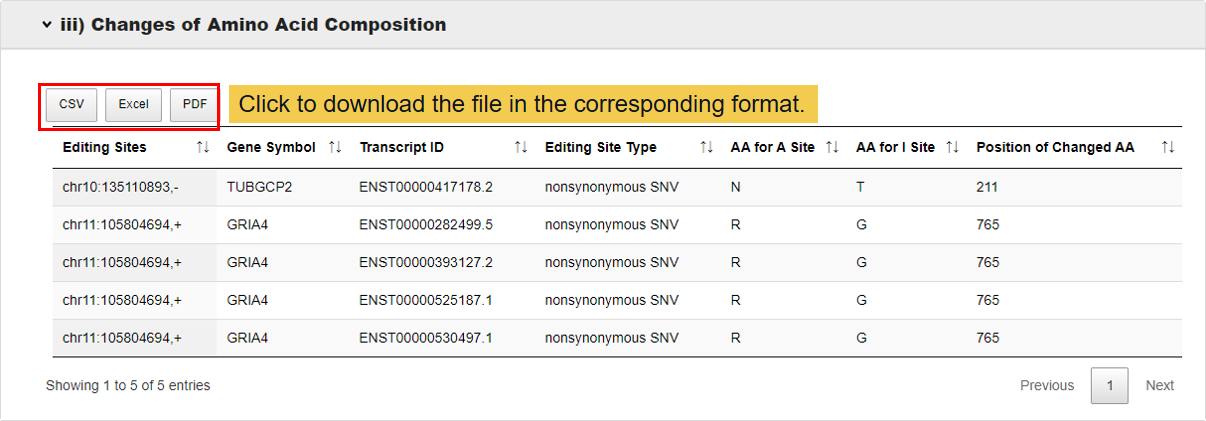
3.3.4 Significantly perturbed biological functions.
This module shows which functions the RNA corresponding to differential editing sites involved in.
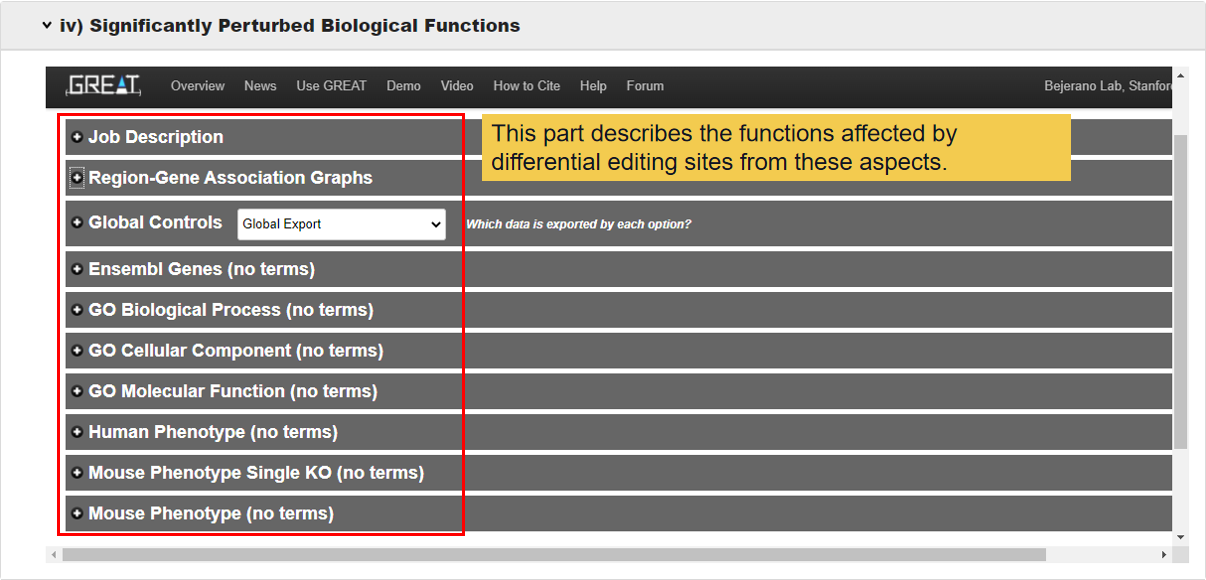
3.4 Pivotal Genes
In this module,genes with the scoring of functional perturbation degree are listed.
RRA Score: RobustRankAggreg score
#FP_Diff_E: Number of differential editing sites which perturbe the functions
#AA: Number of differential editing sites which change amino acid
#miRNA: Number of miRNAs which perturbed by differential editing sites
#RBP: Number of RBPs which perturbed by differential editing sites
#RNA Structure: Number of RNA structures which perturbed by differential editing sites
#Protein Structure: Number of protein structures which perturbed by differential editing sites
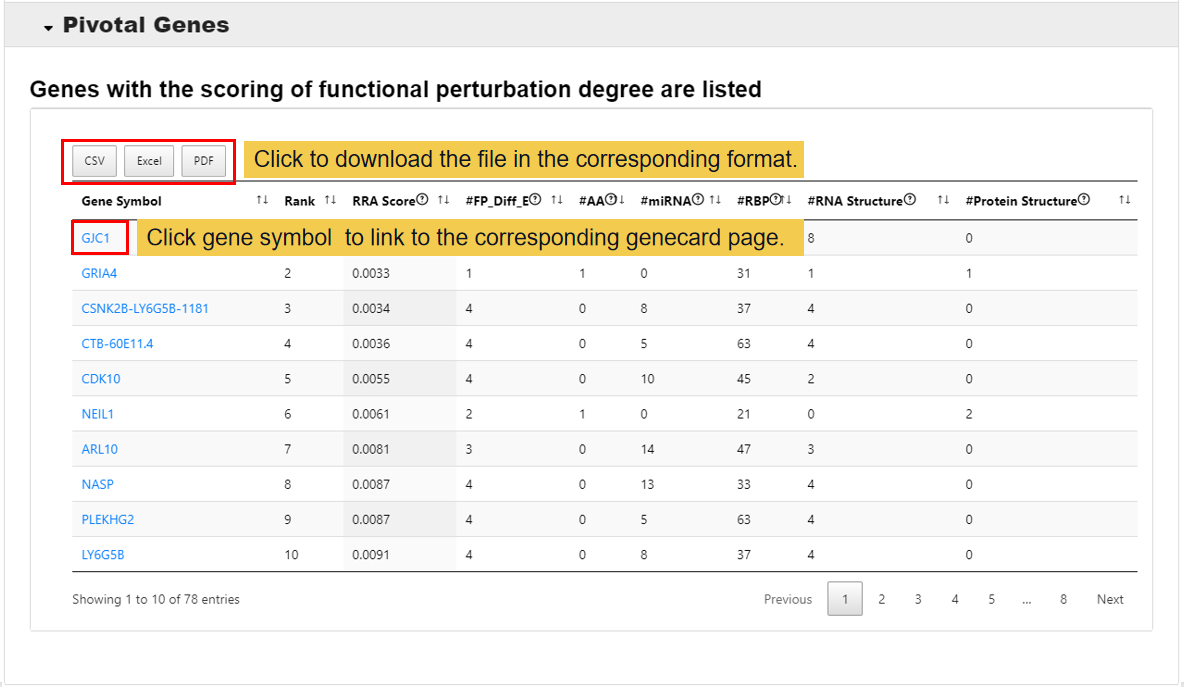
Notes: All figures and tables can download from PRES.
4.Error messages of PRES
A web page error may occur in the following three situation.
4.1 Tasks are only retained for 15 days and then deleted or the Task ID you entered is wrong.
4.2 Your data has been submitted but it hasn’t finished running.
4.3 The format of the file you uploaded is incorrect.