Welcom to using ADRG!
Introduction
The Adverse Drug Reaction related Gene (ADRG) resource is freely available at http://www.bio-bigdata.com/SERT/. It does not require any login or registration and is not password-protected.
ADRG provides a conveniently user-friendly interface to query, browse and visualize detailed information about these adverse drug reactions related proteins. ADRG will be helpful in facilitating the study about the mechanism of ADRs, development of precision medicine and safer drug therapy.
Data sources
The ADR entries under PT level of existing drugs were derived from SIDER database and all ADRs were divided into 26 categories according to ADReCS criteria.The gene entries were derived from NCBI Gene. The approved drug entries were derived from DrugBank and they were classified by ATC. The SNP informations were derived from DrugBank and GWAS Catalog.The risk population, allele frequency and genotype frequency of SNP were derived from 1000 Genomes Project.
Then we apply text mining method to search ADRs, genes and drugs on PubMed (search fileds are title and abstract) from 2006 to 2017 respectively. The search results are purified and filtered, we could get literature about common occurrence of ADR - gene, ADR - drug, gene - drug and ADR - gene - drug.
We not only extracted the relations from literature surveillance of PubMed,but also carried out manual verification.
Tutorial
The ADRP database can be accessed through a simple and user-friendly interface at http://www.bio-bigdata.com/SERT/. Users are allowed to search ADR terms,gene symbols,drug names in the search box of home page.To start a query, users fill out at least one keyword (ADR term, gene symbol, drug name) or three at the same time. When search for an ADR, ADR-gene-drug associations that meet the query criteria are responded in the alphabet order of ADR term along with related genes and drugs. Searching of genes and drugs is in a similar way. Clicking on every ADR-gene-drug association, relevant property information including ADR hierarchy classification, ADR description, WHO code, MedDRA code, synonyms, various gene identifier, protein groups, subcellular location, tissue distribution, drug name, DrugBank ID, drug description will be listed on result page.
Step 1:Input an ADR term(or/and Gene Symbol or/and Drug Name).Then click on the "Submit".(Fig.1)
Step 2:For every ADR-gene-drug association,click on "more" for detailed informations.(Fig.2)
Step 3:The final results page contains detailed ADR informations,gene informations,drug informations,SNP Information From GWAS,reference informations.(Fig.3)
Step 4:For risk population information for each SNP,click on the SNP ID.For every reference,click on PMID for literature.(Fig.3)
Step 5:Risk population information for each SNP.(Fig.4)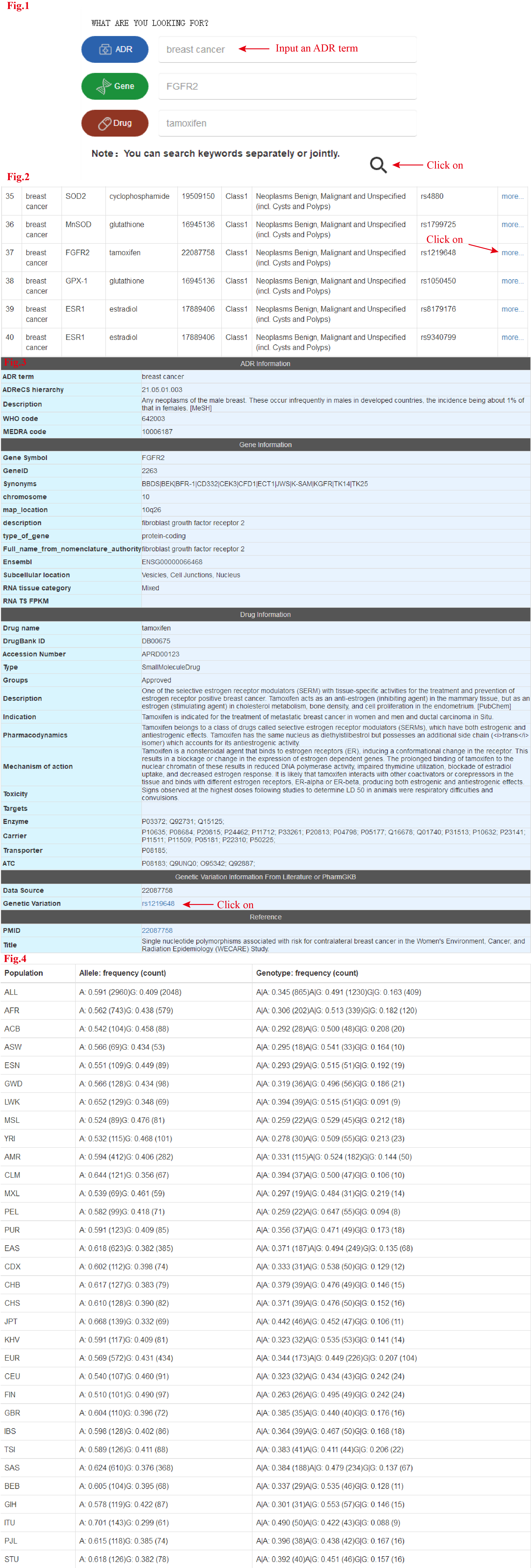
Supplementary explanation
(1) Adverse drug reactions (ADRs):An adverse drug reaction (ADR) is an injury caused by taking a medication.ADRs may occur following a single dose or prolonged administration of a drug or result from the combination of two or more drugs. The meaning of this expression differs from the meaning of "side effect", as this last expression might also imply that the effects can be beneficial.The study of ADRs is the concern of the field known as pharmacovigilance. An adverse drug event (ADE) refers to any injury occurring at the time a drug is used, whether or not it is identified as a cause of the injury. An ADR is a special type of ADE in which a causative relationship can be shown.
(2) Idiosyncratic reaction:Idiosyncratic drug reactions, also known as type B reactions, are drug reactions that occur rarely and unpredictably amongst the population.
(3) Tissue distribution:Tissue-distribution profiles are crucial for understanding the characteristics of cells and tissues in terms of their differential expression of genes.
(4) Subcellular localization:The cells of eukaryotic organisms are elaborately subdivided into functionally-distinct membrane-bound compartments. Some major constituents of eukaryotic cells are: extracellular space, cytoplasm, nucleus, mitochondria, Golgi apparatus, endoplasmic reticulum (ER), peroxisome, vacuoles, cytoskeleton, nucleoplasm, nucleolus, nuclear matrix and ribosomes.Bacteria also have subcellular localizations that can be separated when the cell is fractionated. The most common localizations referred to include the cytoplasm, the cytoplasmic membrane (also referred to as the inner membrane in Gram-negative bacteria), the cell wall (which is usually thicker in Gram-positive bacteria) and the extracellular environment. The cytoplasm, the cytoplasmic membrane and the cell wall are subcellular localizations, whereas the extracellular environment is clearly not. Most Gram-negative bacteria also contain an outer membrane and periplasmic space. Unlike eukaryotes, most bacteria contain no membrane-bound organelles, however there are some exceptions (i.e. magnetosomes).
(5) Adverse Drug Reactions Similiarity Score:
Jaccard index(Tc):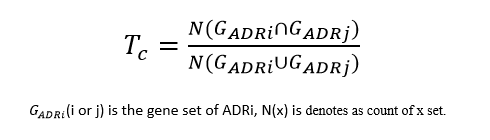
(6) There are 26 different populations which are part of our study from many different locations around the globe. The following table lists these populations and indicates what data we currently have available for them.
| Population Code | Population Description | Super Population Code |
|---|---|---|
| CHB | Han Chinese in Bejing, China | EAS |
| JPT | Japanese in Tokyo, Japan | EAS |
| CHS | Southern Han Chinese | EAS |
| CDX | Chinese Dai in Xishuangbanna, China | EAS |
| KHV | Kinh in Ho Chi Minh City, Vietnam | EAS |
| CEU | Utah Residents (CEPH) with Northern and Western European Ancestry | EUR |
| TSI | Toscani in Italia | EUR |
| FIN | Finnish in Finland | EUR |
| GBR | British in England and Scotland | EUR |
| IBS | Iberian Population in Spain | EUR |
| YRI | Yoruba in Ibadan, Nigeria | AFR |
| LWK | Luhya in Webuye, Kenya | AFR |
| GWD | Gambian in Western Divisions in the Gambia | AFR |
| MSL | Mende in Sierra Leone | AFR |
| ESN | Esan in Nigeria | AFR |
| ASW | Americans of African Ancestry in SW USA | AFR |
| ACB | African Caribbeans in Barbados | AFR |
| MXL | Mexican Ancestry from Los Angeles USA | AMR |
| PUR | Puerto Ricans from Puerto Rico | AMR |
| CLM | Colombians from Medellin, Colombia | AMR |
| PEL | Peruvians from Lima, Peru | AMR |
| GIH | Gujarati Indian from Houston, Texas | SAS |
| PJL | Punjabi from Lahore, Pakistan | SAS |
| BEB | Bengali from Bangladesh | SAS |
| STU | Sri Lankan Tamil from the UK | SAS |
| ITU | Indian Telugu from the UK | SAS |
These populations have been divided into 5 super populations
- AFR, African
- AMR, Ad Mixed American
- EAS, East Asian
- EUR, European
- SAS, South Asian
When the code ALL is used this means that all individuals from that release are being considered.
