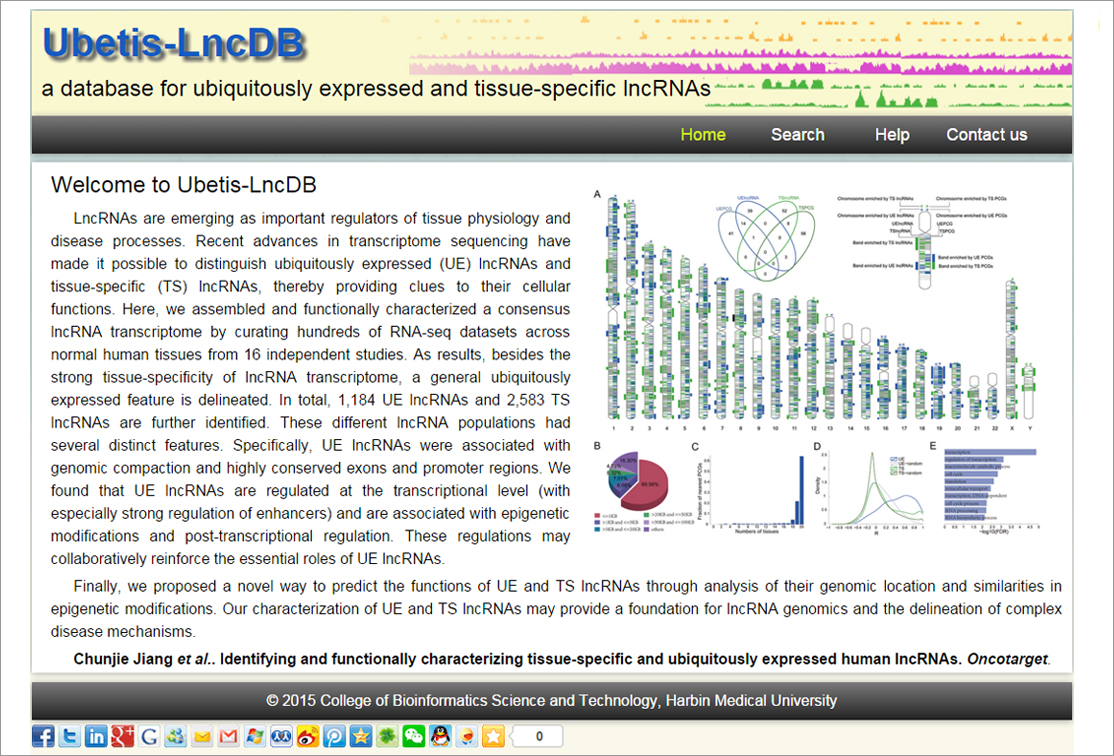
Recent advances in transcriptome sequencing have made it
possible to distinguish ubiquitously expressed long non-coding
RNAs (UE lncRNAs) from tissue-specific lncRNAs (TS lncRNAs),
thereby providing clues to their cellular functions. Here, we
assembled and functionally characterized a consensus lncRNA
transcriptome by curating hundreds of RNA-seq datasets across
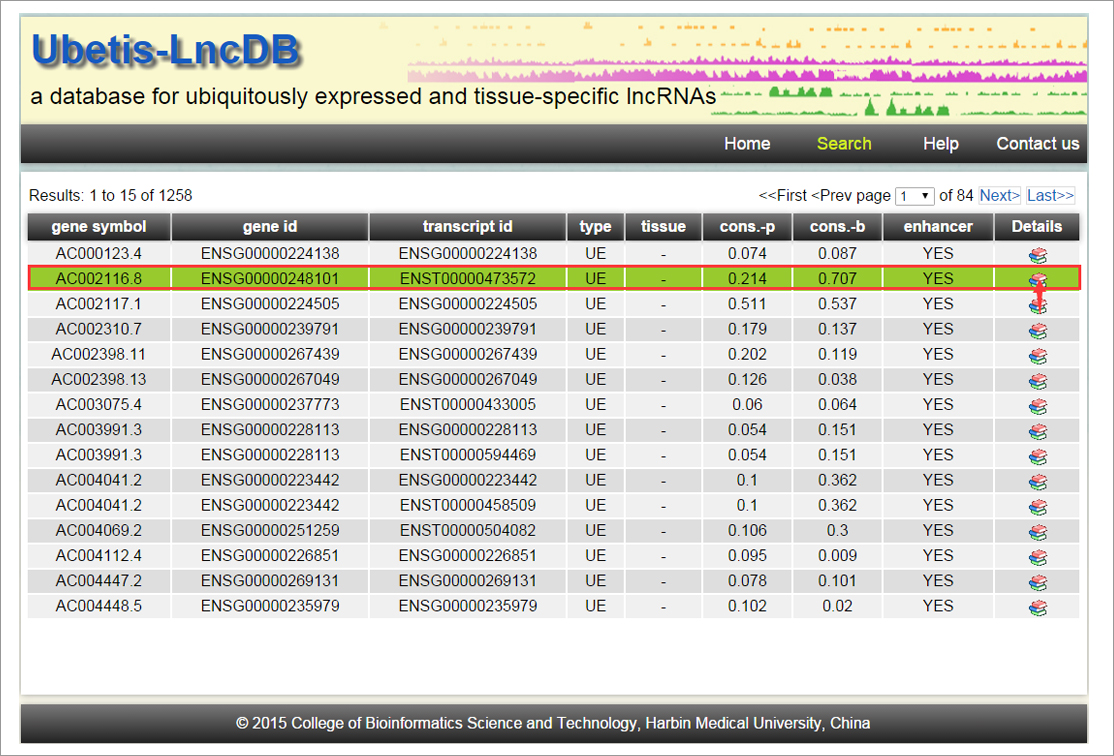
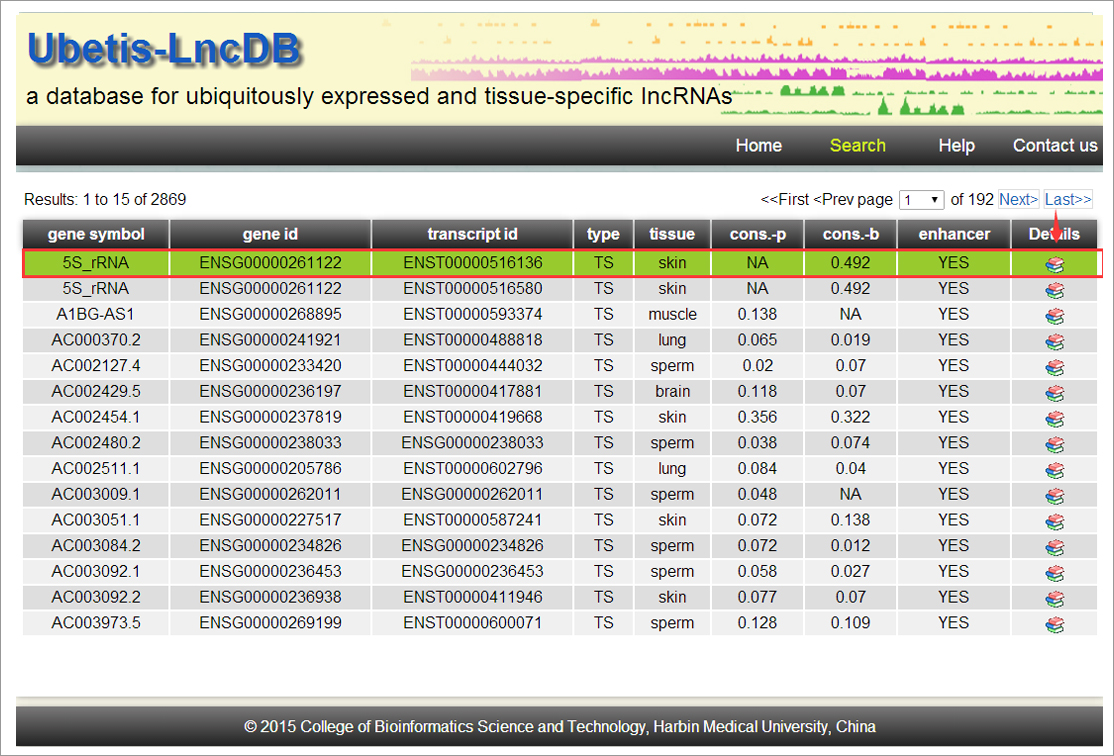
normal human tissues from 16 independent studies. In total,
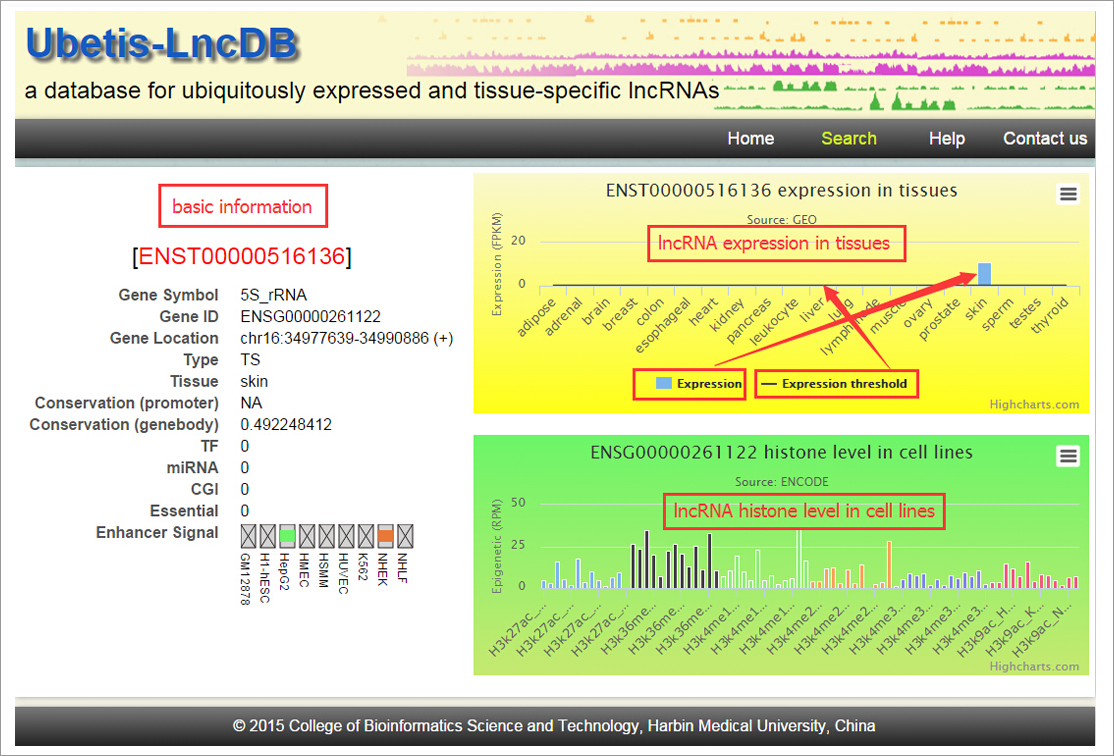
1,184 UE and 2,583 TS lncRNAs were identified. These different
lncRNA populations had several distinct features. Specifically,
UE lncRNAs were associated with genomic compaction and highly
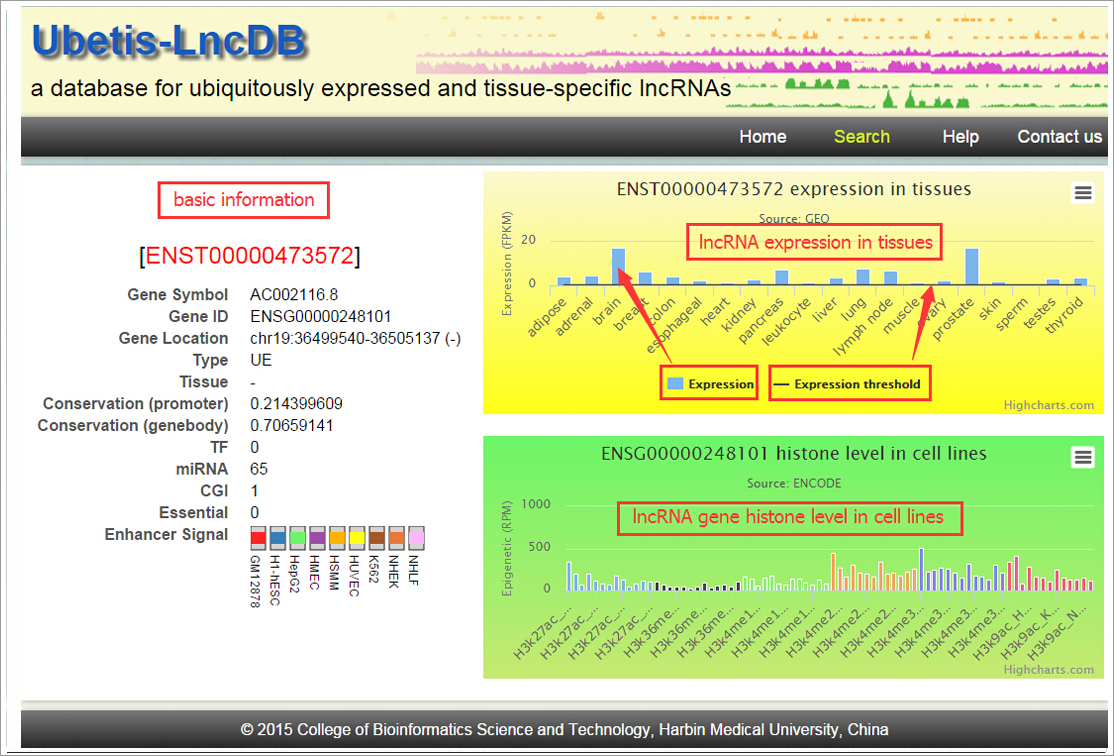
conserved exons and promoter regions. We found that UE lncRNAs
are regulated at the transcriptional level (with especially
strong regulation of enhancers) and are associated with
epigenetic modifications and post-transcriptional regulation.
Based on these observations we propose a novel way to predict the functions of UE and TS lncRNAs through analysis of their genomic location and similarities in epigenetic modifications. Our characterization of UE and TS lncRNAs may provide a foundation for lncRNA genomics and the delineation of complex disease mechanisms.
 .
.