CLING (Cancer LncRNA Identification via Network Grouping) is a computational framework which is developed for human cancer lncRNA prioritization. It is an intuitive and accessible tool that allow users with limited bioinformatic skills to rapidly access and visualize data pertinent to their research. Currently, CLING can be applied to optimize lncRNAs in 16 types of human cancer in whole genome (lncRNA annotation file: GENCODE V19).
The main idea of CLING is to present an overall prioritization list of candidate lncRNAs through fusing the multiple separate prioritization results that are generated from different lncRNA-centric networks. This framework involves 3 main steps:
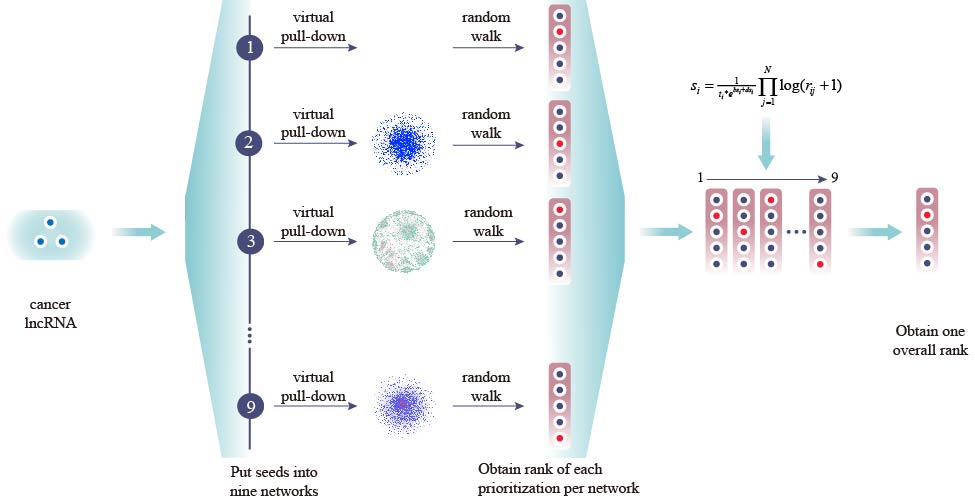
Figure 1. Workflow of the CLING
Firstly, a list of known cancer lncRNAs (seed lncRNAs) were retrieved from Lnc2Cancer database, and virtually put into corresponding networks as the source nodes.
Secondly, candidate lncRNAs were ranked based on the score obtained from random walk with restart (RWR)algorithm with the advancement of source nodes, which results in a single prioritization list for each network.
Finally, data fusing algorithm (MdDI) was used to integrate all separate prioritization results and lncRNA topological properties to form an overall prioritization list.
The "Prioritization" page allows users to input a list or submit a text file that contains the candidate lncRNAs, and then returns a result page, depicting the overall optimization results, the separate optimization results in each network and the sub-network of candidate lncRNA.
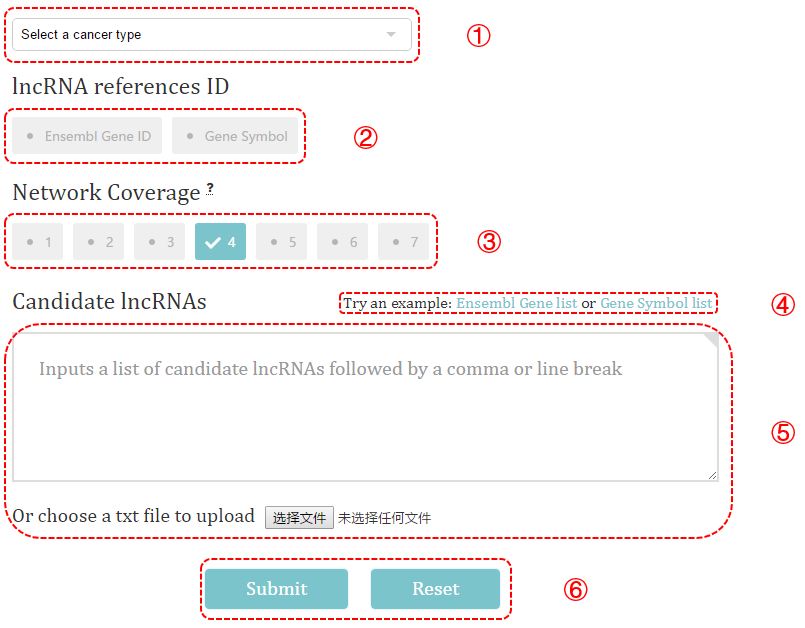
Figure 2. Data submission
① Select a cancer type and view the seed lncRNAs used in prioritization.
② Choose the lncRNA reference ID of the entered lncRNAs. Here, two types of lncRNA ID are allowed, which are Ensembl gene ID and gene symbol.
③ Select the network coverage, an option that allowed users to selectively prioritize cancer lncRNA. For instance, the default value 4 means only the lncRNAs involved in at least 4 networks will be optimized.
④ Use the example we provided to test the performance of CLING.
⑤ Insert a list of lncRNAs into the text box or submit a text file.
input format: [lncRNA][,][lncRNA] or [lncRNA][\n][lncRNA]
⑥ Click Submit button and wait for a while to see the result page.
The returned result page contains a summary of the submission information and optimization result (Figure 3) and a detail table (Figure 4) describing the ranking and sub-network of each candidate lncRNA.
Summary
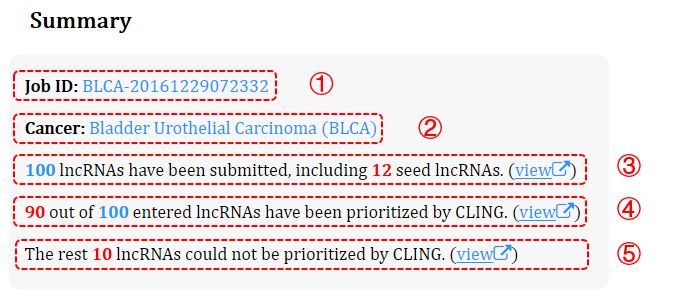
Figure 3. Summary of the optimization
① Job ID.
② Recheck the selected cancer type.
③ The number of the submitted lncRNAs (nonredundant) and the seeds their contained. Clicking this link (view) will direct to details of the contained seeds.
④ The number of the lncRNAs that are optimized by CLING. Clicking this link (view) to view the details.
⑤ The number of the lncRNAs that are not optimized by CLING. Clicking this link (view) to view the details.
Detail
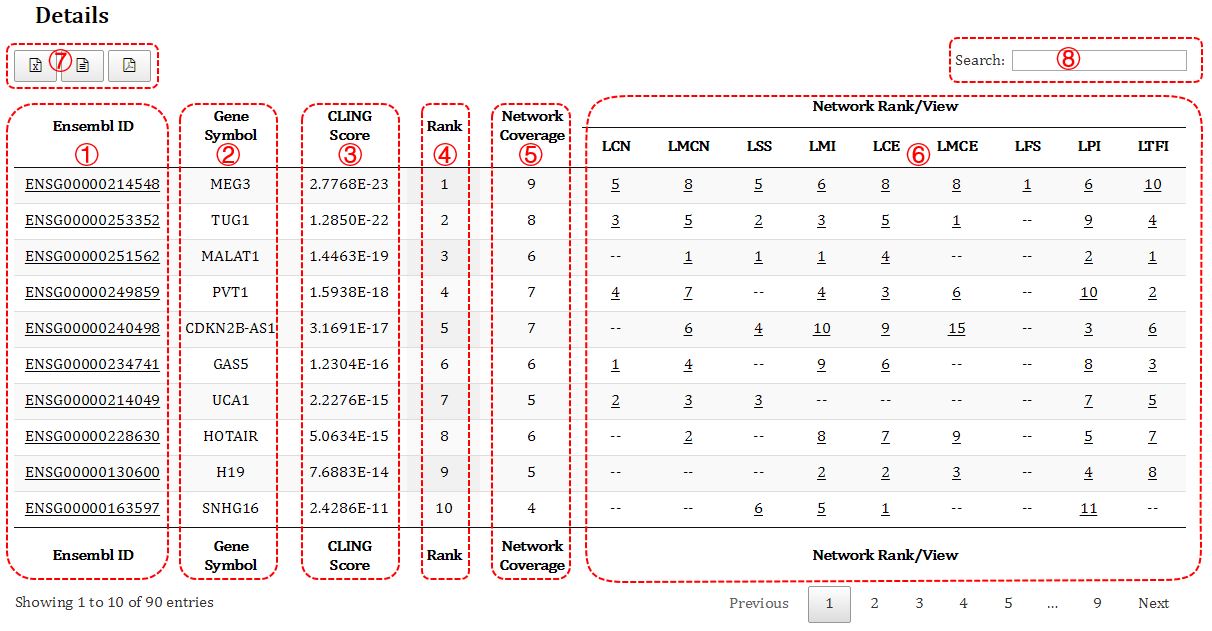
Figure 4. Detail result of the optimization
① The Ensembl gene ID of the optimized lncRNAs.
② The gene symbol of the optimized lncRNAs.
③ The overall score calculated by CLING. The higher ranking, the lower score.
④ The overall ranking of the candidate lncRNAs.
⑤ The number of the network that an candidate lncRNA involved in. For instance, 8 means this lncRNA appears in 8 networks.
⑥ The ranking of the candidate lncRNAs that are optimized by an individual network, where "--" means the candidate lncRNA is missing in corresponding network. Clicking this number (ranking) will direct to subnetwork of the candidate genes which formed by its direct neighbors in the network (Figure 5).
⑦ Buttons to retrieve the prioritization result. Currently, 3 types of file (excel, csv, pdf) are provided for users to download.
⑧ The search box allows users to quickly retrieve the prioritization result of one particular lncRNA.
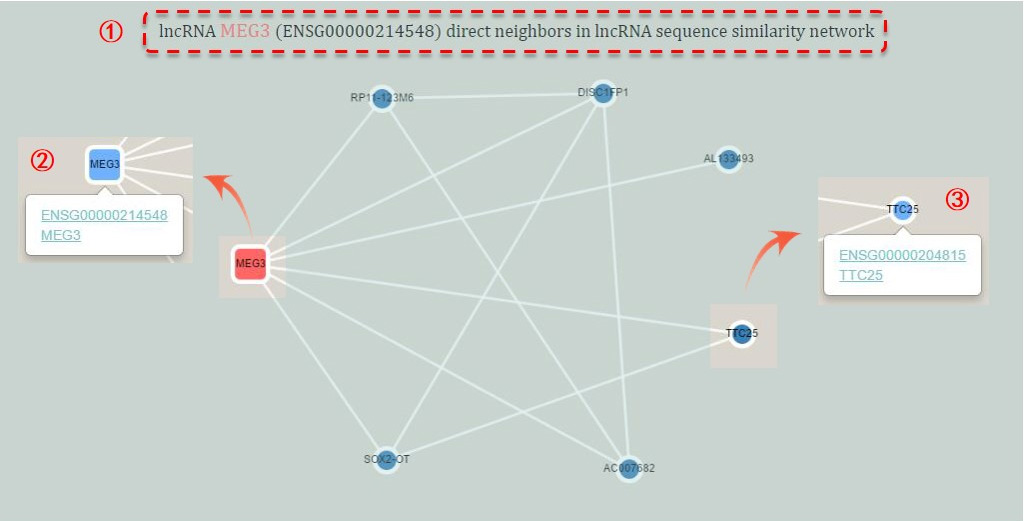
Figure 5. Sub-network of an indivdual network
① The name of the corresponding lncRNA and sub-network.
② The core lncRNA is marked by rounded rectangle node. The other nodes are presented as circle nodes. And the selected node will be highlighted by sky blue. An example of MEG3 is on the left.
③ By clicking a node, you can see the additional information and some links under the node. Click to view more about the node.
Query help
The "Query" page allows users to input different key words (lncRNA or cancer name) to query related data prioritized by CLING. If the entered term is lncRNA name (Ensembl ID or gene symbol), the results page will return the ranking information of this lncRNA in different cancer types (Figure 6); while the top 200 optimized lncRNAs will be returned if the entered key word is cancer name (Figure 7).
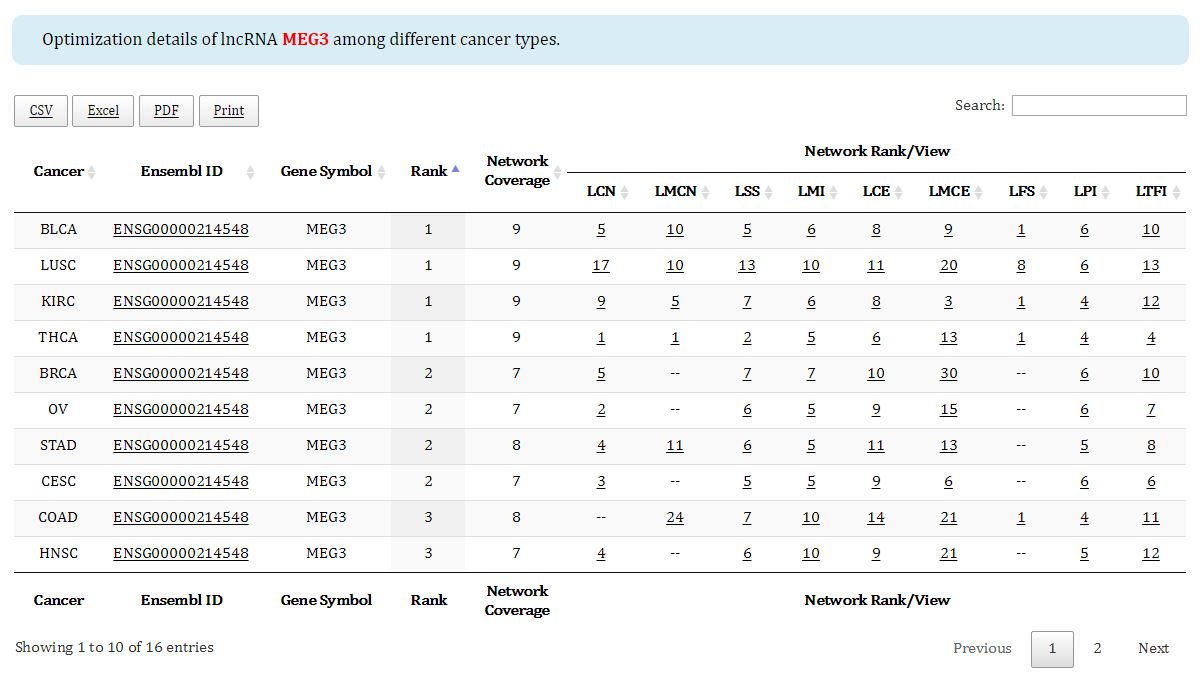
Figure 6. Prioritization result of an individual lncRNA among different cancers
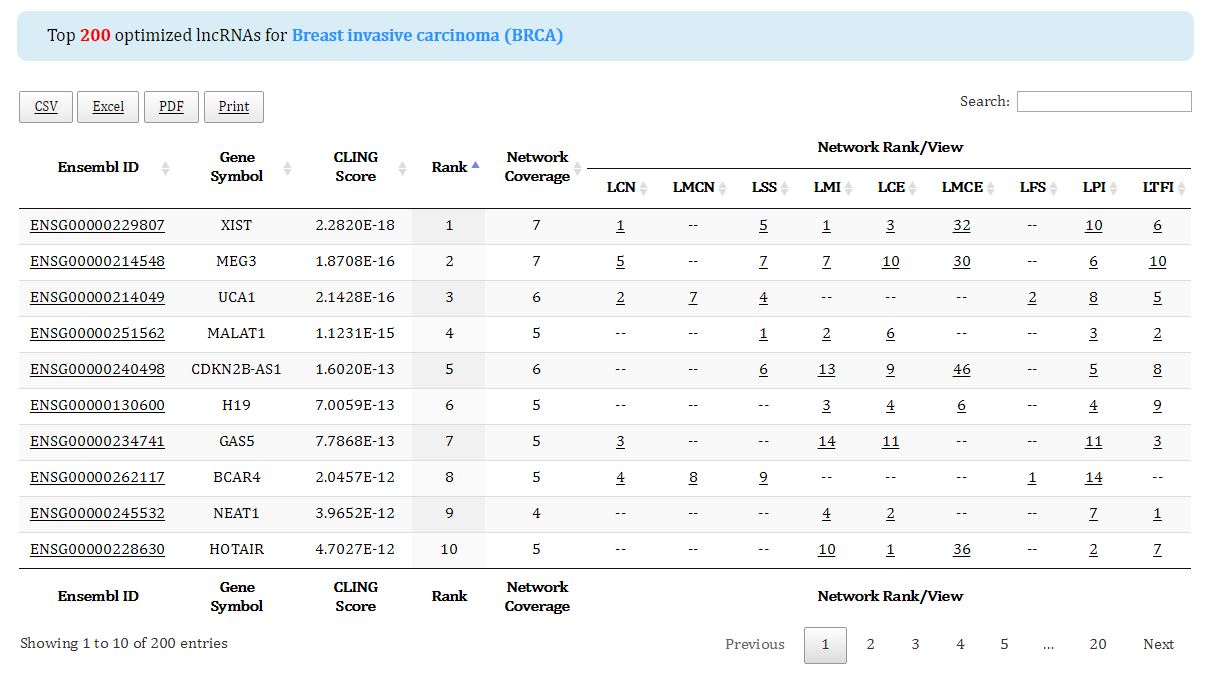
Figure 7. Top 200 optimized lncRNAs of the queried cancer
For the comprehensive prioritization list, please move to the "Download" page.
Download help
The "Download" page allows users to download the overall prioritization list predicted by CLING as well as the experimentally supported cancer lncRNAs used as seed lncRNAs (Figure 8).
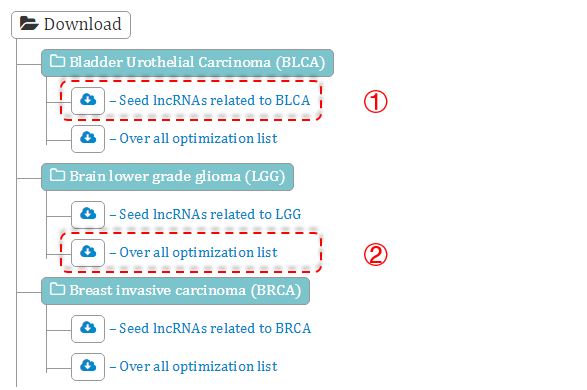

Figure 8. Download page
① Experimentally supported cancer-related lncRNAs.
② The overall prioritization result of a cancer type predicted by CLING based on the seed lncRNAs.