Help |
Welcome to MERIT! Here is a step to step introduction about how to use the web to search the RBP, gene and mutation in cancer or search the correlation between RBP and Target Gene.

MERIT: Mutational Effect on RNA Interactome Topology
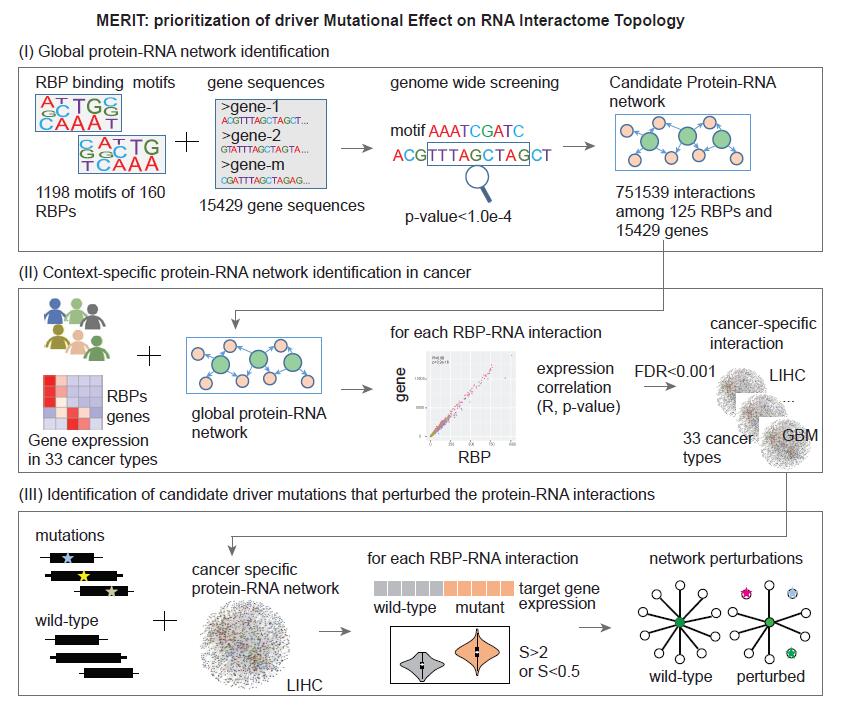
Here, we proposed a three-step computational method (MERIT) to identify the candidate driver mutations that perturbed the protein-RNA interaction network in cancer. Step-1: Firstly, the 1,350 genes coding for RNA binding proteins analyzed in this study were obtained from a recent study (1), including those with high confidence for RNA binding and genes annotated as RNA binding in Ensembl. To identify the genome-wide RBP-gene regulation in cancer, we firstly retrieved the 1,198 motifs of 160 human RBPs from ATtRACT database (2). In addition, genome-wide sequences of 15,429 genes were obtained from one of the previous studies (3). The FIMO program was used to search these sequences for occurrences of known RBP motifs (4). Motifs with Benjamini and Hochberg adjusted p-values less than 1.0e-4 were regarded as binding sites in the gene. Step-2: Emerging evidence indicated that if genes were regulated by RBPs, their expression levels should be correlated (5,6). Thus, we integrated genome-wide expression of RBP and genes to investigate the context-specific RBP-gene regulation in each cancer types. Here, we explored the regulation via spearman correlation method. For each pair of RBP-gene predicted above, we computed the spearman correlation coefficient (R) as follow: R(x,y)=cov(RX,RY)/(σRX*σRY) =E[(RX-μRX)(RY-μRy)]/(σRX*σRY) =(E[(RX-μRX)(RY-μRy)])/(√(E[〖RX〗^2 ]-〖[E[RX]]〗^2 ) √(E[RY^2 ]-〖[E[RY]]〗^2 )) Where RX was the rank of expression for RBP x in samples and RY was the rank of expression for gene y in samples. μRX and μRY were the mean rank of RBP x and gene y. σRX and σRY were the standard deviation of RX and RY. E was the expectation. All the candidate RBP-gene pairs with P-adjusted<0.01 were identified as RBP-gene regulation in each cancer type. Step-3: To identify the somatic mutation mediated RBP-gene interactions, we started with a hypothesis that mutations located in the RNA binding sites were likely to disrupt the RBP binding. Disruption of RBP binding might affect the expression of target genes. Based on the expression correlation in each cancer type, we divided the RBP-gene pairs into two groups: one contained the RBPs promoting the expression of target genes, and the other contained RBPs inhibiting the expression of target genes. Next, we divided the cancer samples based on whether the sample had a specific mutation. The expression levels of target gene k were compared between the mutated group and the normal group and the expression alteration score (S) was calculated as follow: S(k)=M ̅_k/W ̅_k =(∑_(i=1)^m▒〖x_i/m〗)/(∑_(j=1)^w▒〖x_i/w〗) Where the m and w is the number of mutated samples and wild-type samples. If the expression levels of target genes showed 2-fold reduction (S<0.5) in the mutated group and the RBP promoted the expression of the target genes in normal samples, we defined these pairs as decreased activation. In contrast, if the expression levels of target genes showed 2-fold increase (S>2) in the mutated group and the RBP inhibited the expression of the target genes in normal samples, we defined these pairs as decreased inhibition. All the RBP-target gene pairs were assembled as mutation-mediated RBP-gene interaction rewiring.
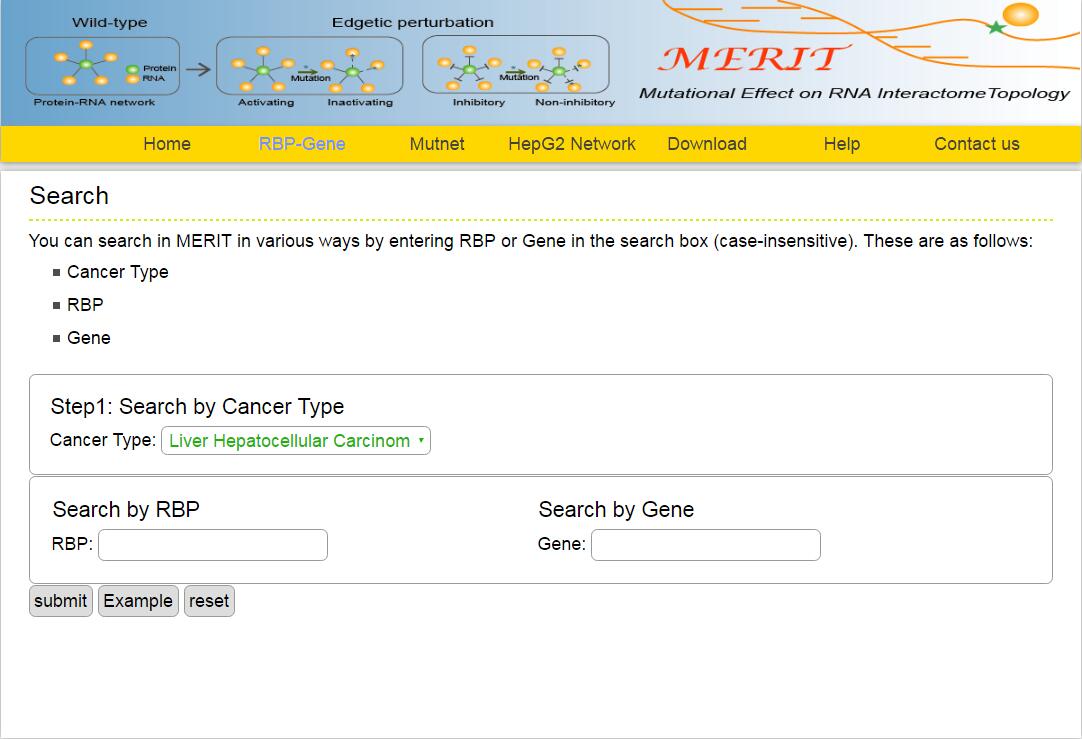
The MERIT provides an interface to retrieve the information about the RBP-Gene in cancer. The current release of MERIT mainly contains the results in 33 kinds of cancer. Users can search by individual RBP or by individual gene to view the RBP-Gene association in a certain cancer. Here, we provide an example where users can click on "examples" and "submit", to check out the information about association.
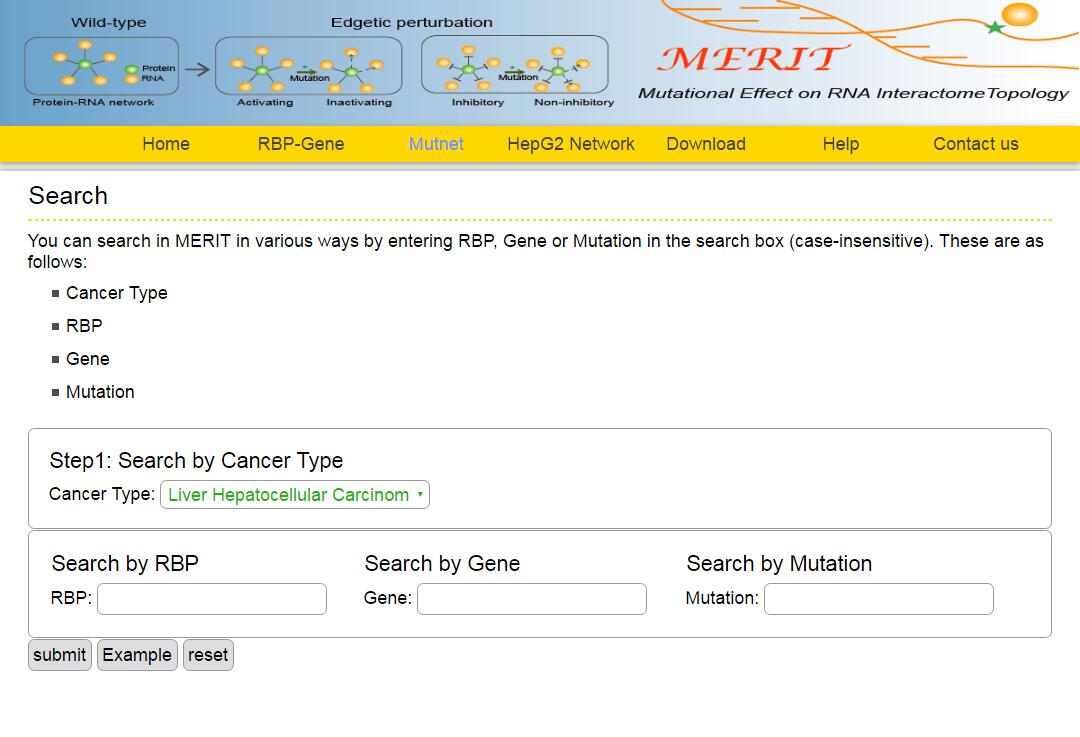
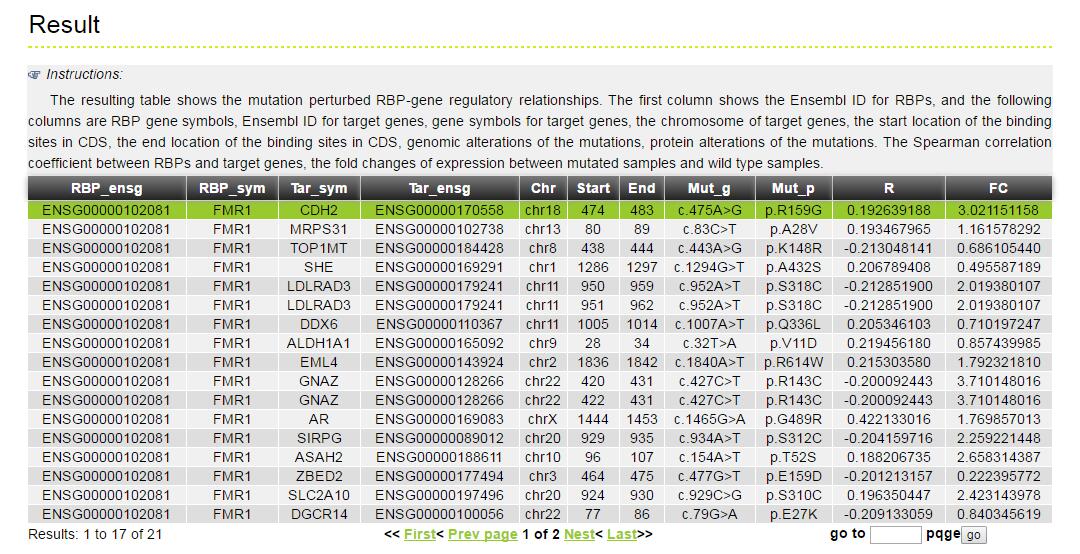
The MERIT provides an interface to retrieve the information about the RBP-Mutation-Gene triplets in cancer. The current release of MERIT mainly contains the results in 28 kinds of cancer. Users can search by individual RBP or by mutation or by individual gene to view the RBP-Mutation-Gene association in a certain cancer. Here, we provide an example where users can click on "examples" and "submit", to check out the information about triplets.
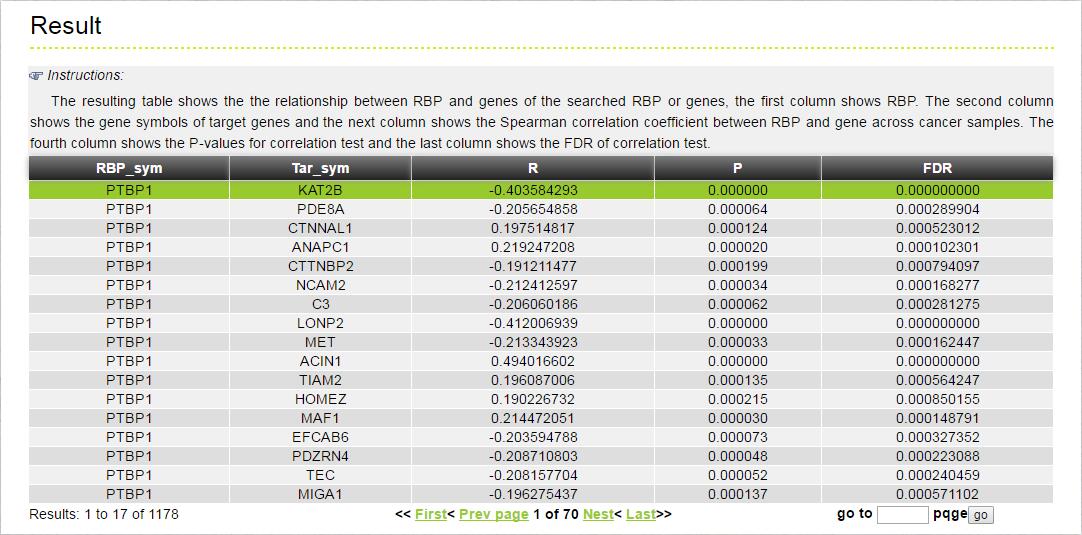
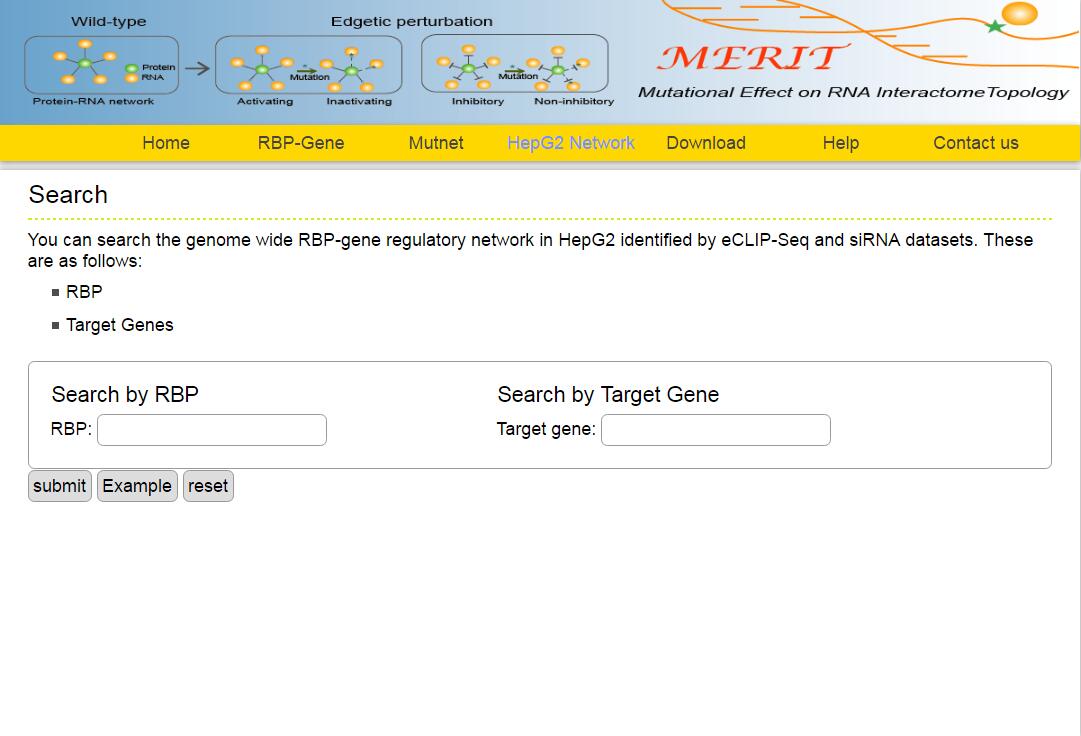
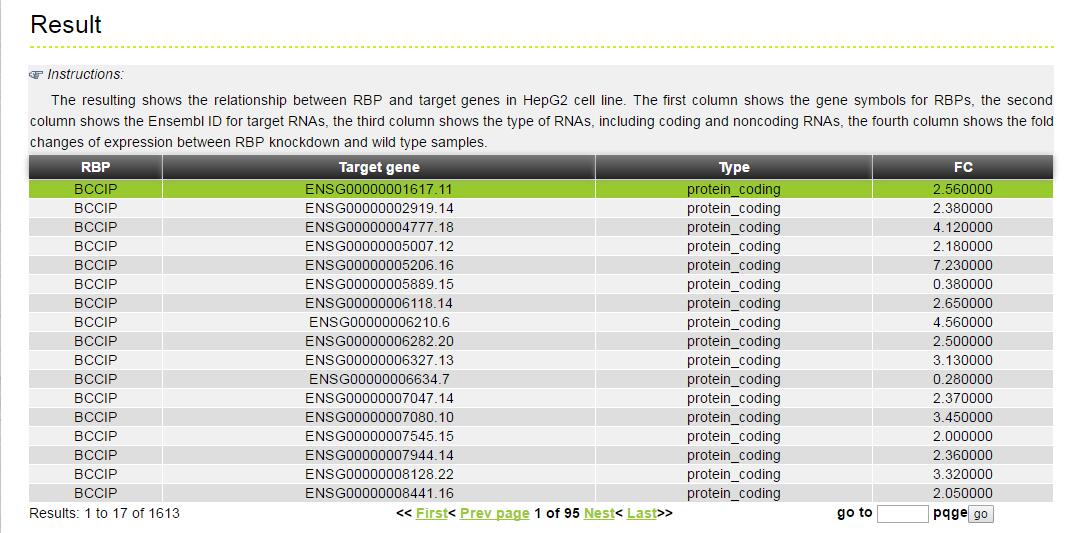
The MERIT provides an interface to retrieve the information about the RBP-(Target Genes). Users can search by individual RBP or by target gene to view the RBP-(Target Gene) association. Here, we provide an example where users can click on "examples" and "submit", to check out the information about association.
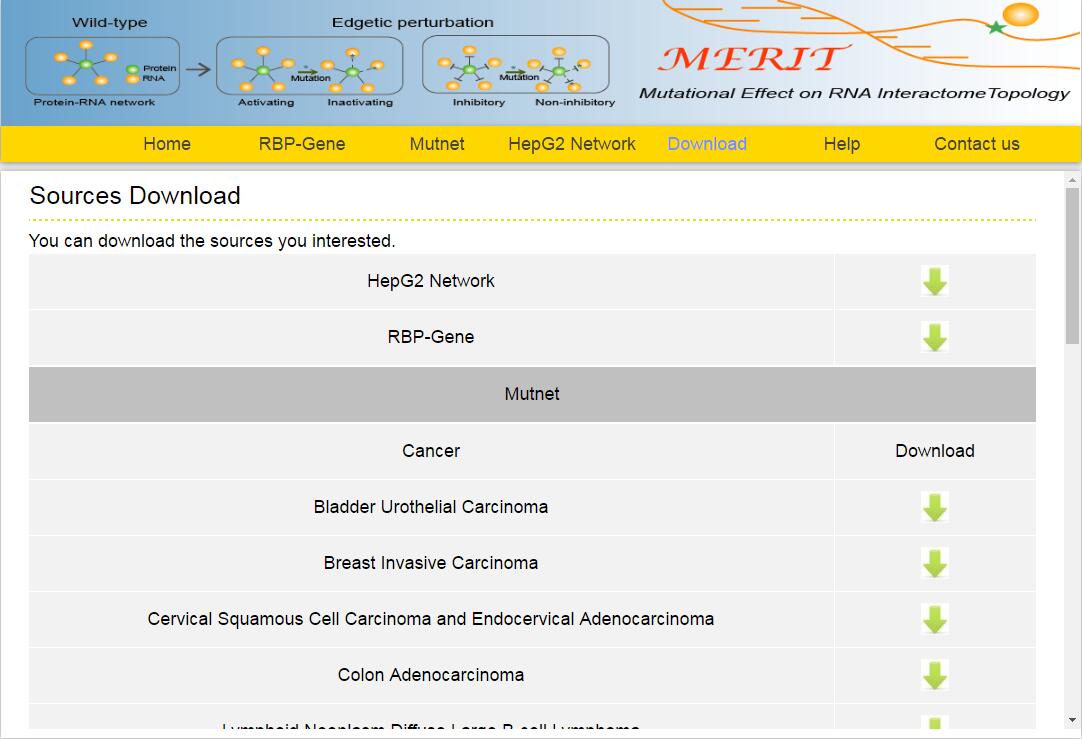
In addition, download links are provided, which allowed you to batch download the information you interested in.