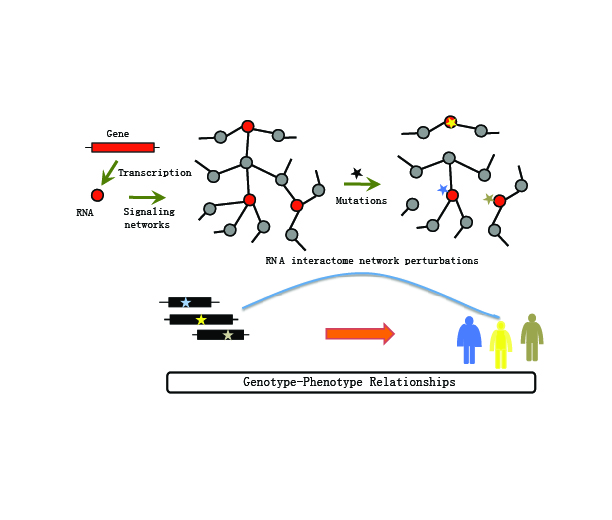
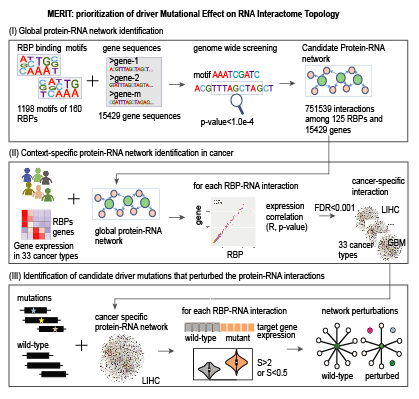
Welcome to MERITRNA binding proteins (RBPs) play important roles in regulating gene expression. However, decoding the genome wide RBP regulatory network as well how cancer related mutations impair the RBP regulatory activities in the context of network remain mostly undetermined. Recently, we explored the genetic alteration pattern of 1,350 RBPs and found that widespread deleterious mutations are likely to occur at the surface of RBPs across 33 types of cancer, suggesting RBP regulatory network perturbations. Thus, we constructed genome-wide RBP-regulatory network by integration of sequence binding screen and expression correlation. RBP regulatory network analysis highlighted synergetic regulation among RBPs. In addition, we found that mutations selectively targeted functional important genes (cancer genes, core fitness genes or conserved genes) and perturbed the RBP-gene regulatory network in cancer. Perturbed genes were mainly involved in cell adhesion related functions. Comparison of the mutation perturbed RBP regulatory profile, we observed that different mutations leading to different RBP regulation perturbation often result in distinct cancer types. All these regulatory patterns were further validated in independent datasets. Finally, we also analyzed RBP-lincRNA regulation and revealed lincRNA-LIMT involved in DNA repair and immune response related functions, which might be a candidate noncoding biomarker in cancer. Taken together, our results provided a valuable resource for characterizing mutation perturbed RBP regulatory network in cancer, paving the way to insights into the genotype-phenotype relationships underlying cancer. This resource provided the genome-wide RBP-gene regulatory network across 33 types of cancer and HePG2 cell line. The users can easily query the interesting RBP, gene and somatic mutations in specific cancer types. |
Statistics:• Cancer type: 33 • Cancer samples: 10,489 • Mutations analyzed: 2,571,512 • 1,198 motifs of 160 RBPs; 15,429 ORF sequences • 136 eCLIP experiments for 68 RBPs • 450 shRNA-Seq experiments for 225 RBPs |