Home
MG-ncExplorer is a comprehensive resource for exploring ncRNA involvement in microglia. On the home page, you can see its workflow, data statistics, and interaction network.
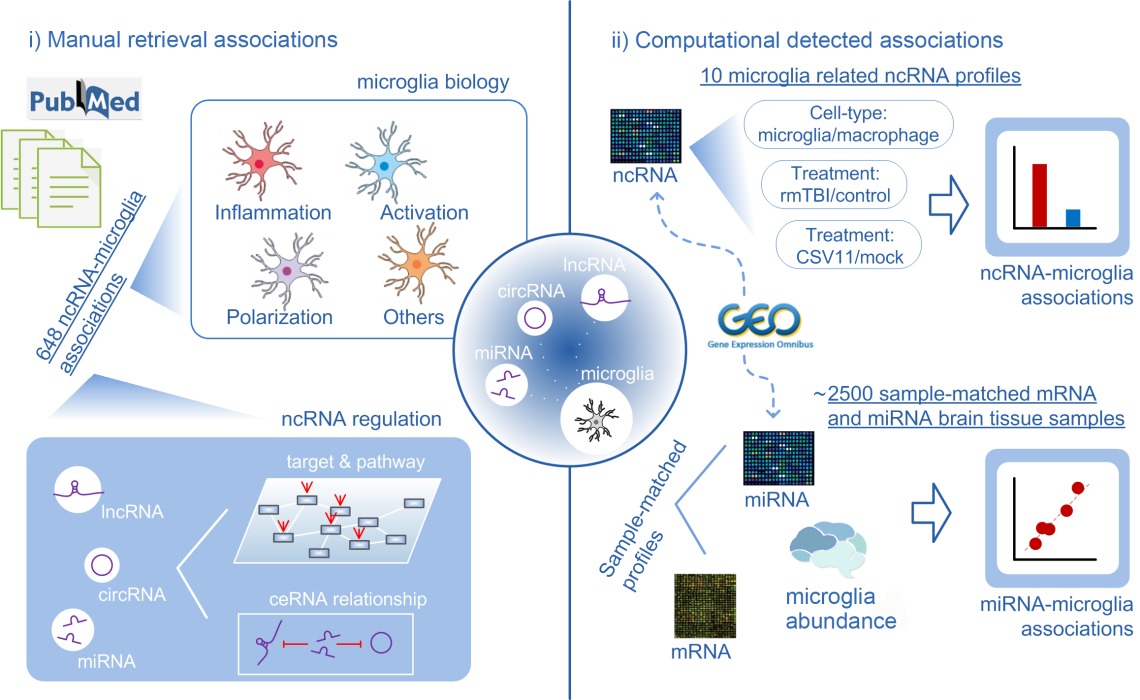
Workflow
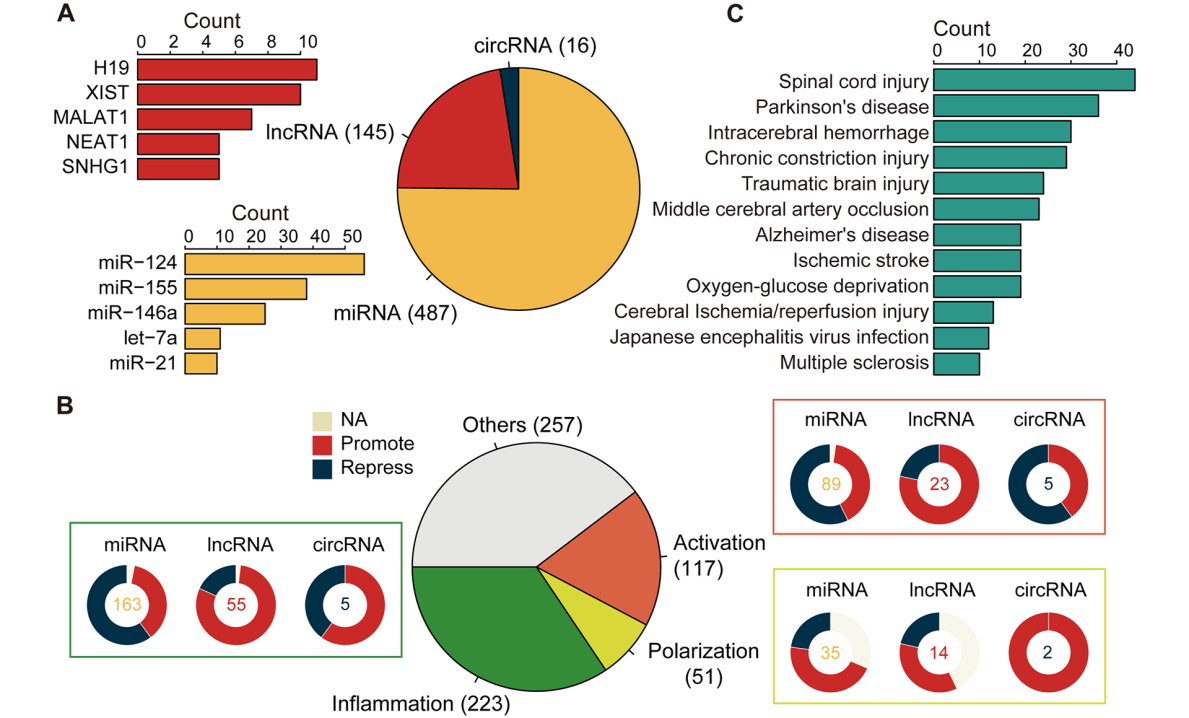
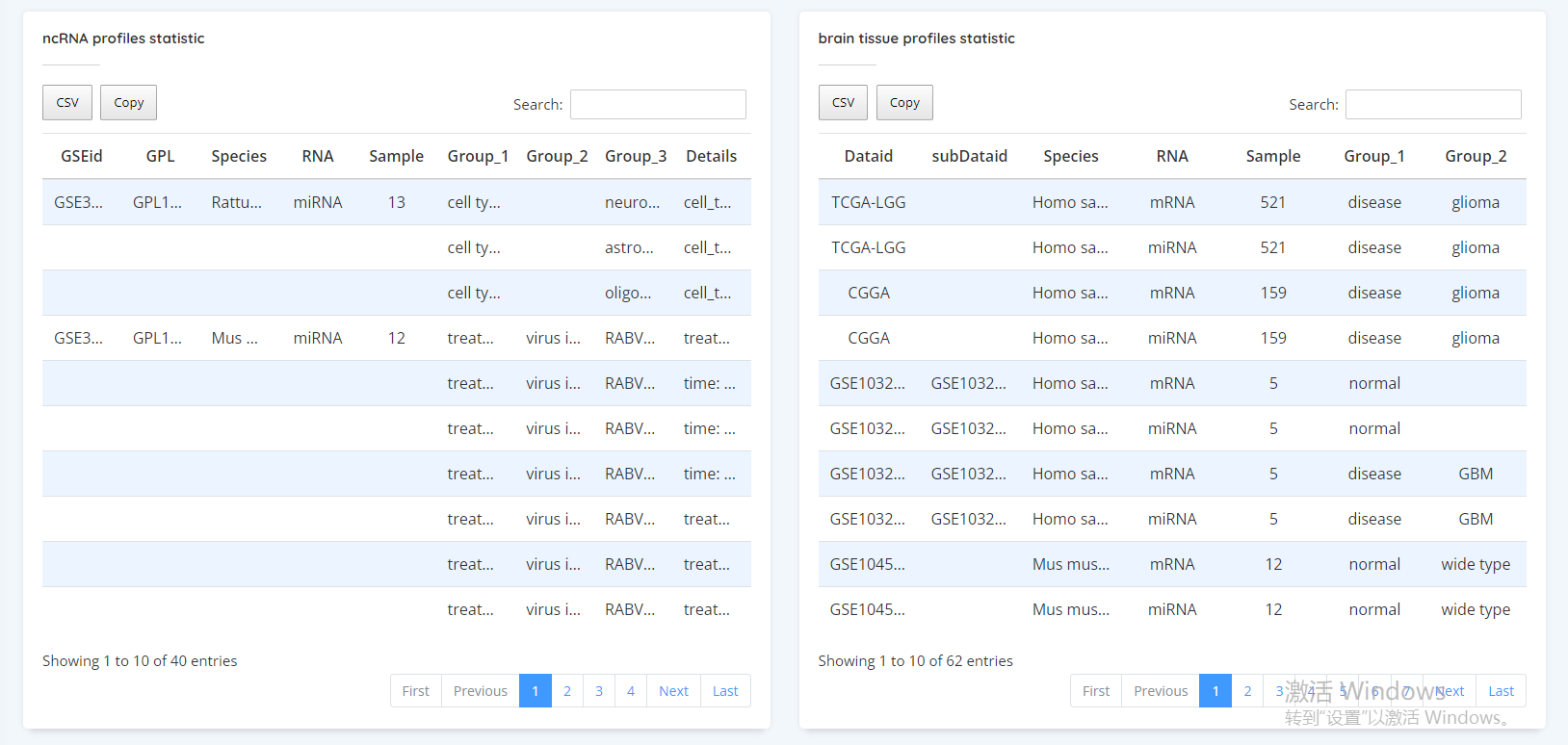
Data Statistics
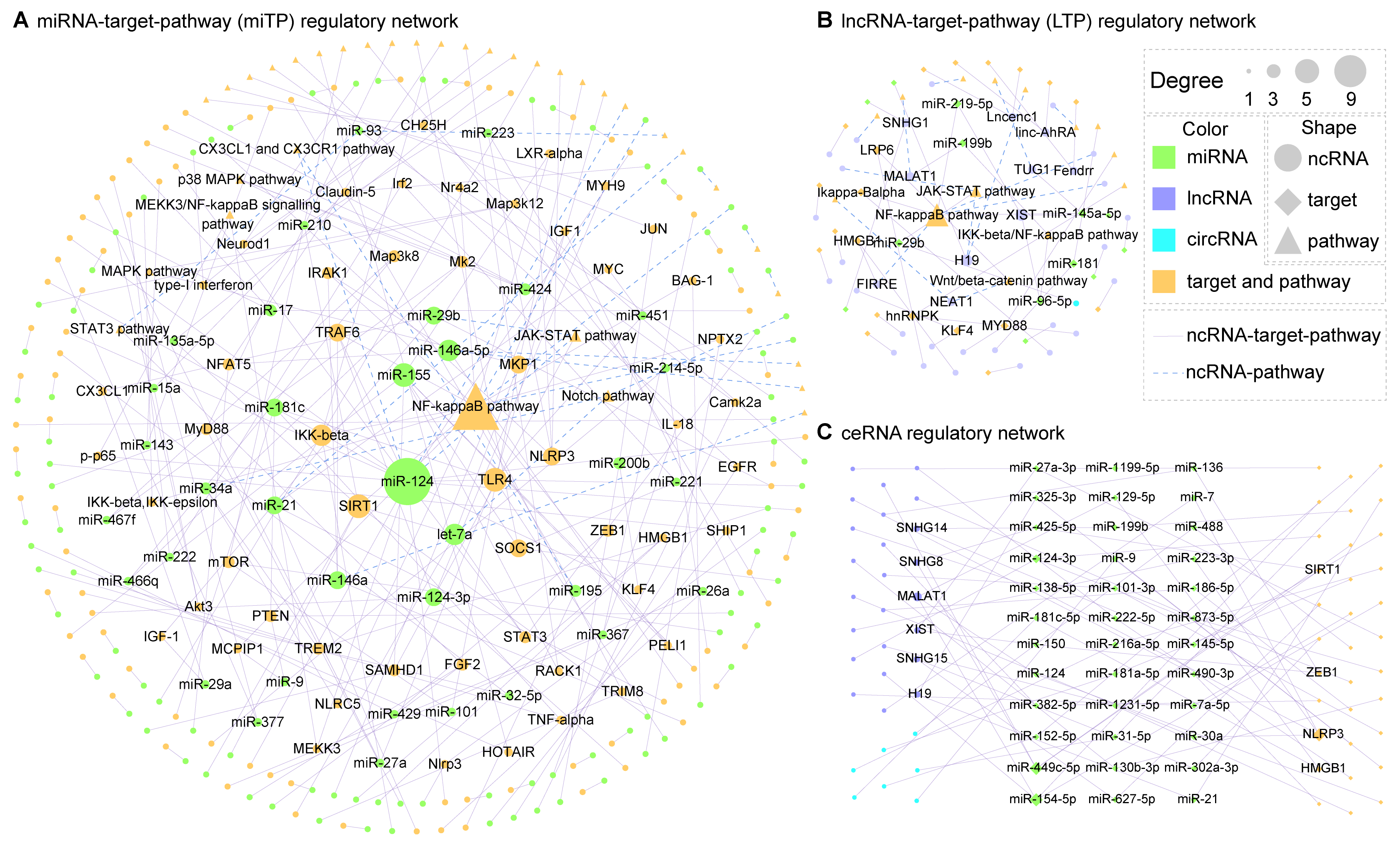
Interaction Network
Manual retrieval
In this section, users can view the relationship between 648 pairs of microglia and ncRNA (including miRNAs, lncRNAs and circRNAs) from about 500 articles by searching or browsing.
Search
Users can enter the ncRNA or regulatory target or pathway they interested in the search bar on the left to view the details containing these keywords.
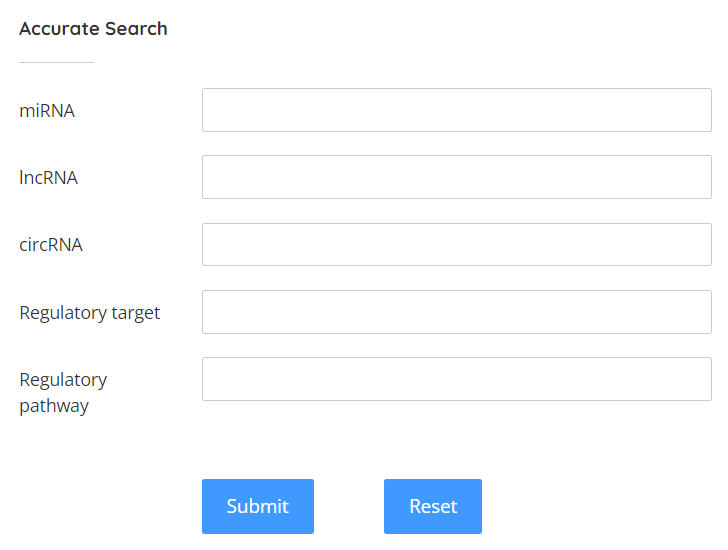
Browse
In addition to searching, you can also browse all data according to ncRNA type, microglia type and model by clicking on the tree folder on the browsing page.
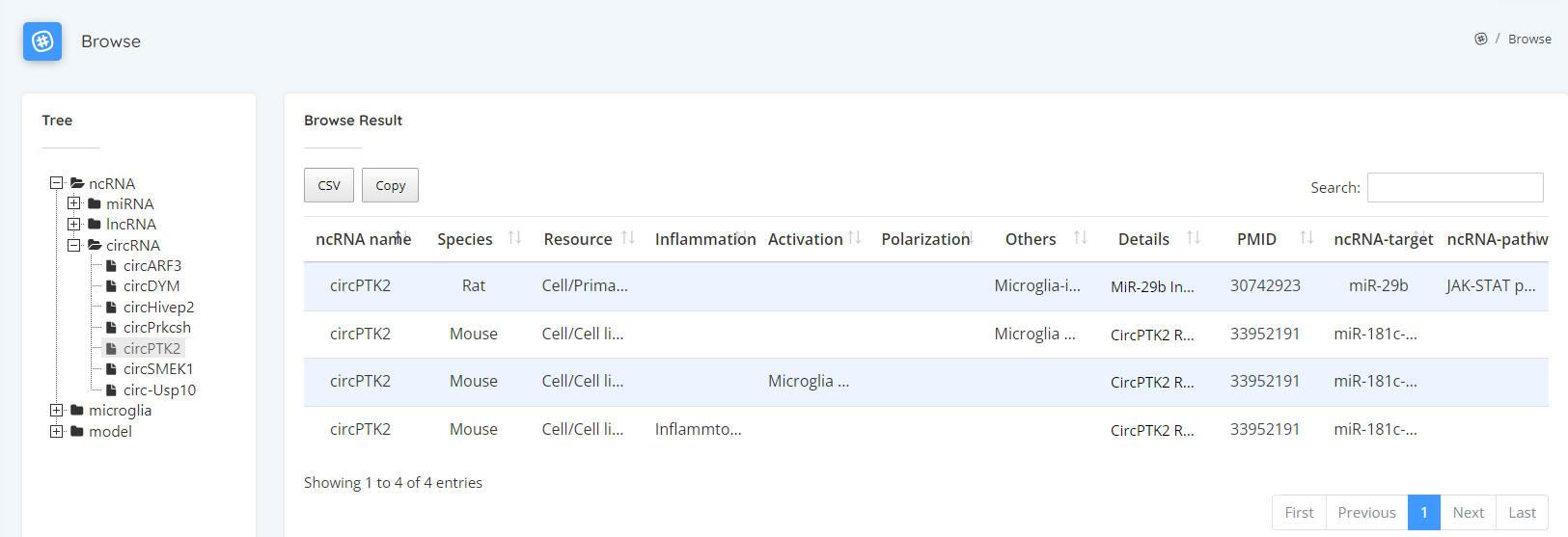
Computational detection
In this section, we collected a large number of ncRNA data and predicted the ncRNA relationship in microglia through high-throughput sequencing, which is divided into the following two functional modules:
Cell & microglia
In this section, we do the expression differences of ncRNA in microglia under different states (cell type, treatment, disease, sex) at the cellular level. After selecting the options you want to do difference, click the submit button to get the heat map, volcano map and list of differentially expressed genes.
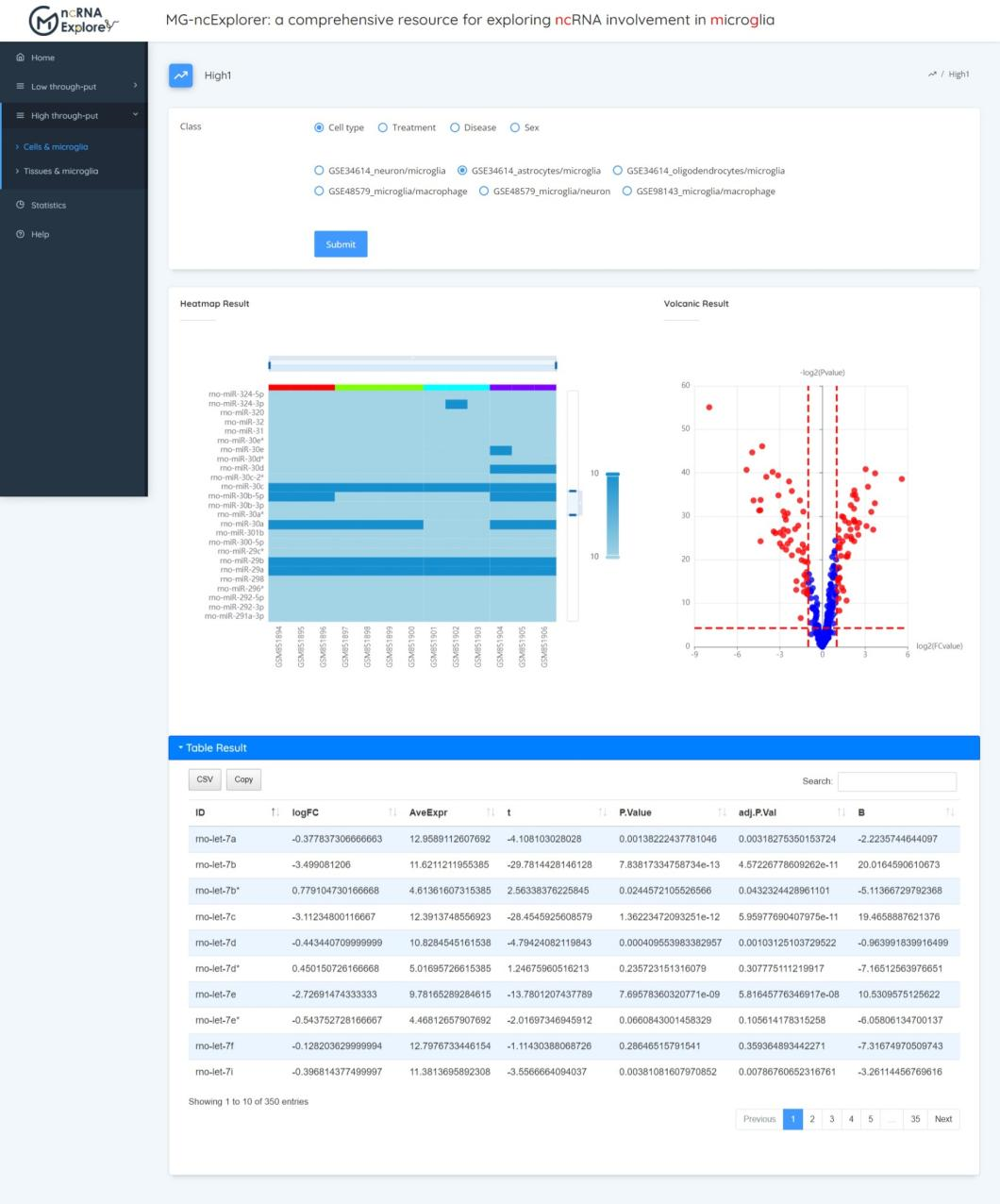
Tissue & microglia
In this section, we estimate the proportion of microgliain the tissue (degree of infiltration) according to the marker of microglia. Then the correlation between microglia infiltration and miRNA expression was calculated. Users can first select disease or normal, and then select the data set of interest to view the correlation between microglia infiltration and ncRNA expression value.
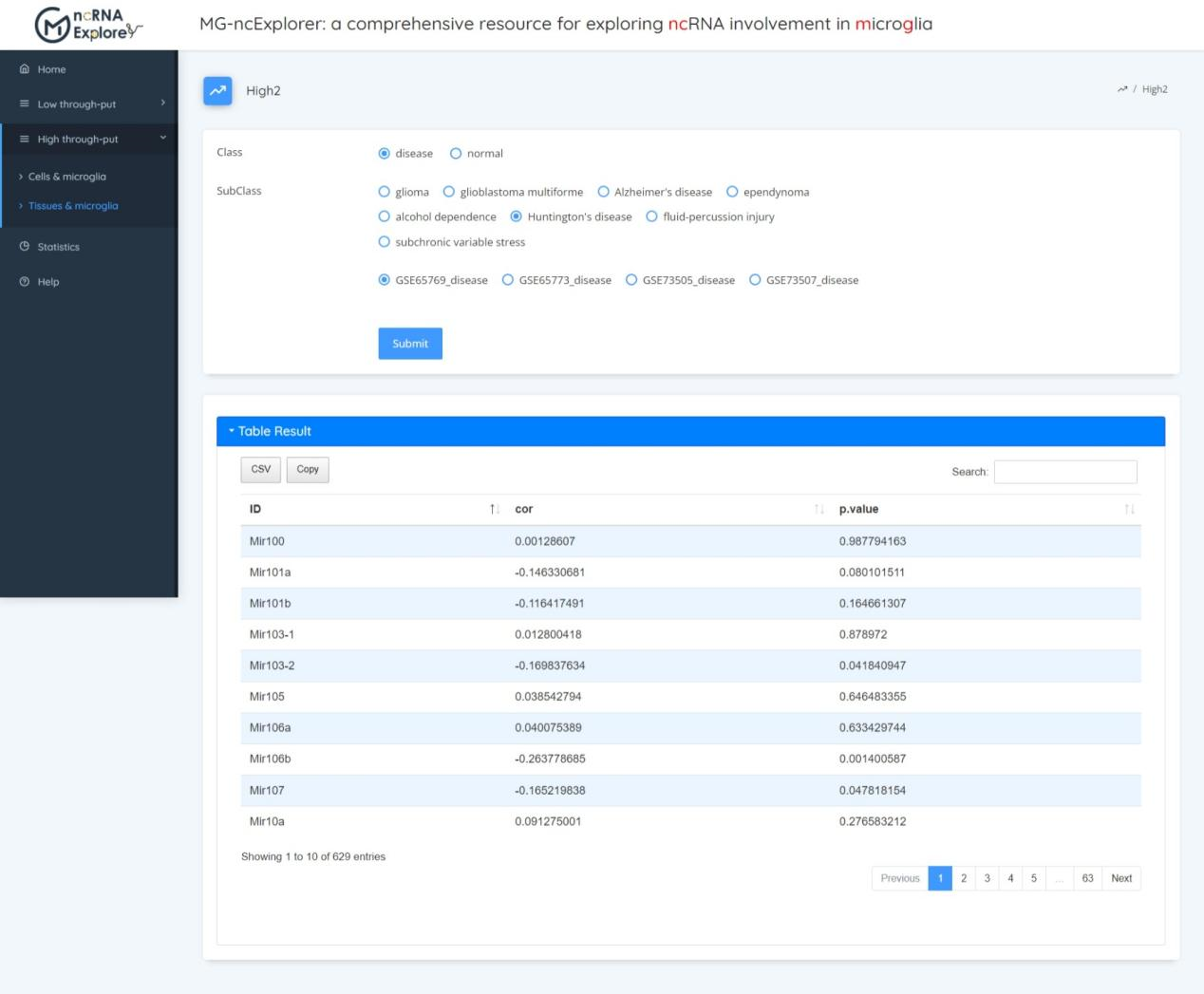
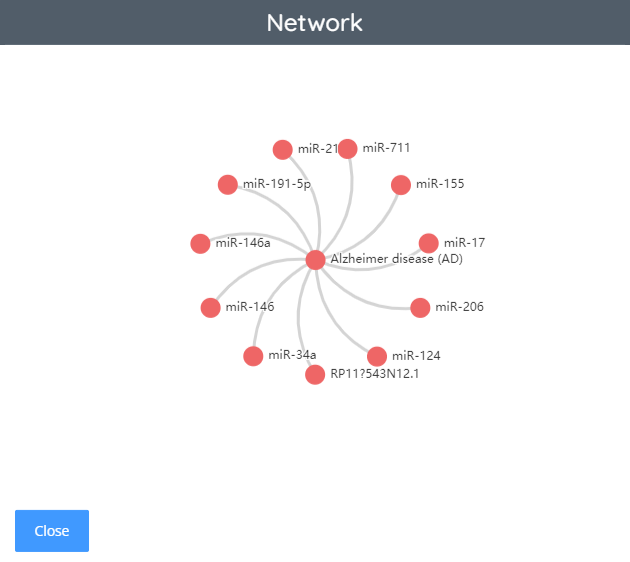
Statistics
In the statistics section, we can view some relevant statistical information of ncRNA in this database, including pathways, disease models and so on. This section also shows the network diagram, which users can click to view according to their own needs
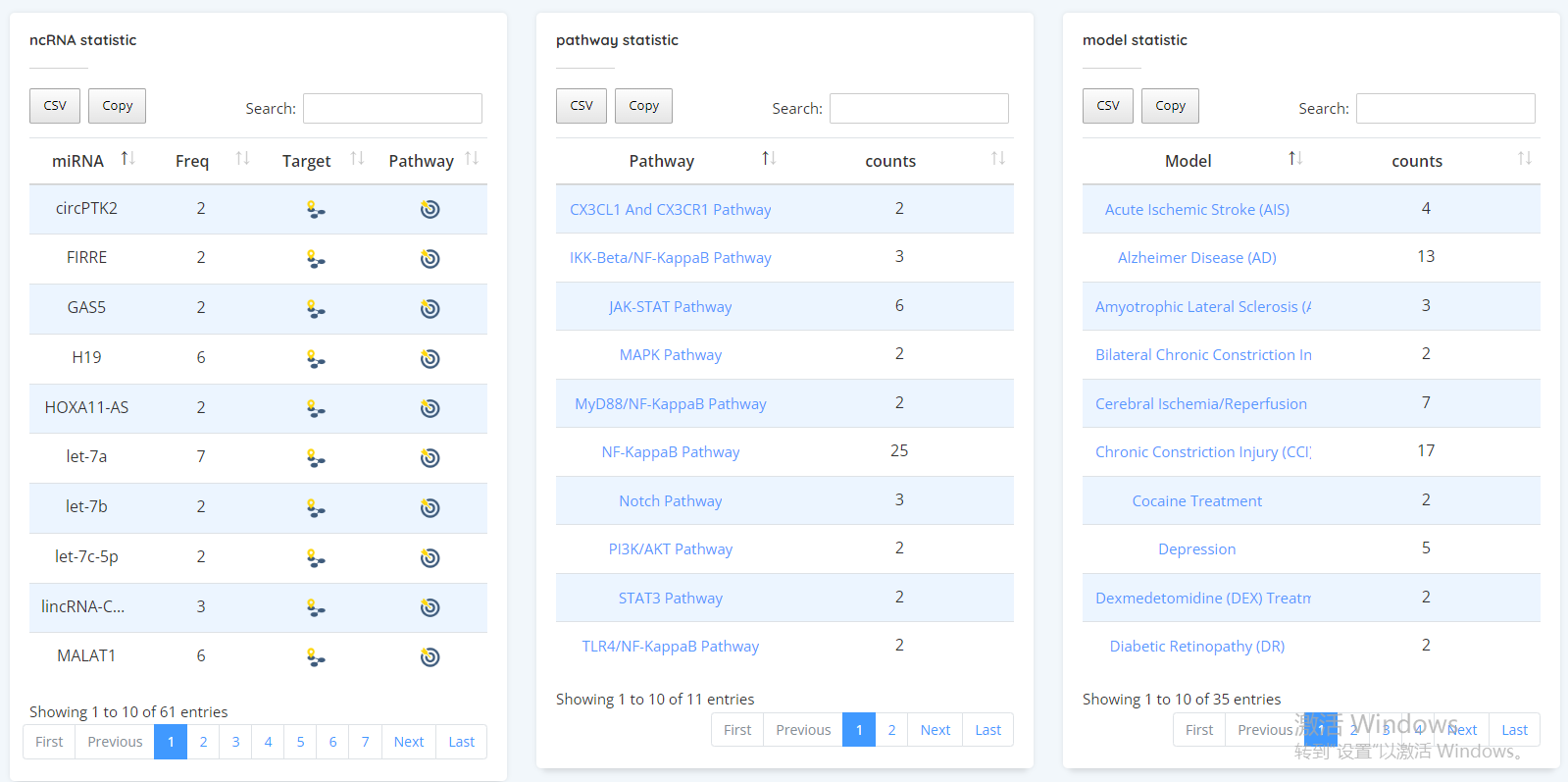
miR-124 target network
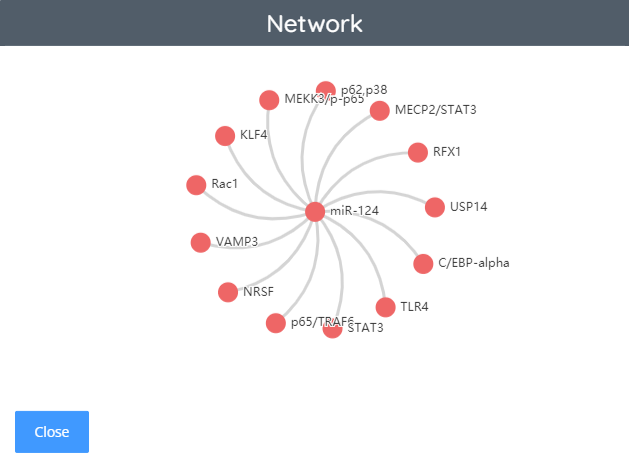
JAK-STAT pathway - ncRNA network
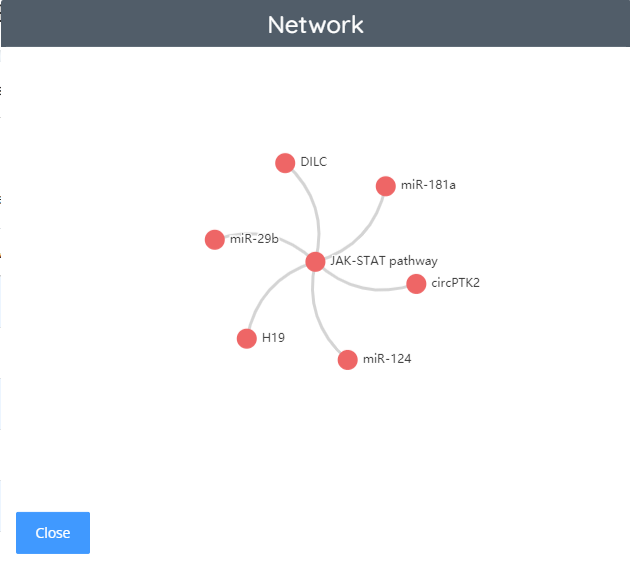
Alzheimer’s disease - ncRNA network
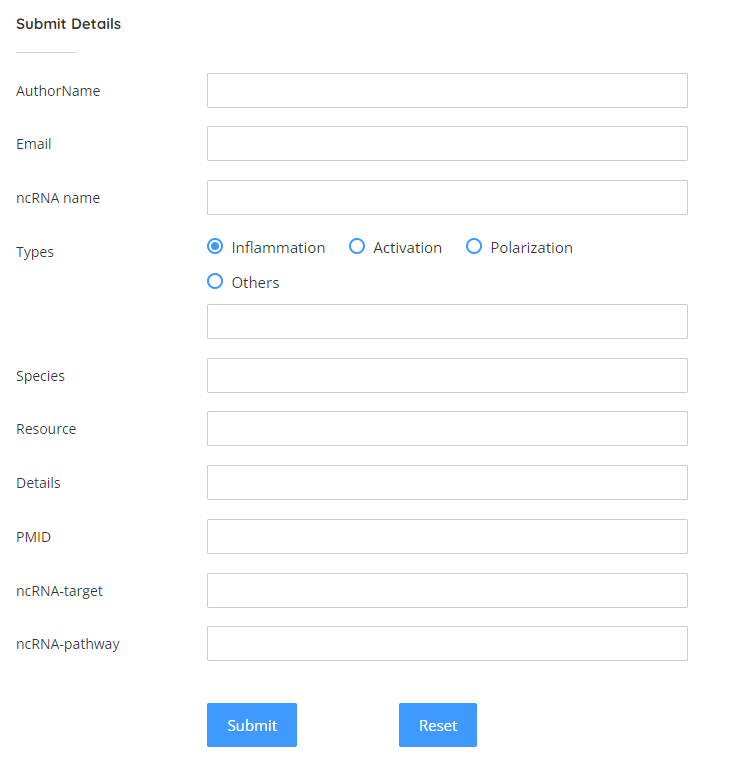
Submit
In the meanwhile, we welcome users to submit the latest ncRNA information related to microglia. Here you shall submit the author’s name, email address, ncRNA name, regulation type, species, source and other information we need.