CPAD overview
CPAD houses a comprehensive manually curated collection of experimentally supported associations between human cancers and biological pathways and their upstream regulators (microRNA, lncRNA, TF, drug, etc.). The CPAD database also integrates multi-omics and clinical data for nearly 10,000 patients across 28 cancer types from TCGA project and drug related data from DrugBank and CellMiner databases. In CPAD, these data are directly linked to each cancer pathway and CPAD provides a flexible platform to explore the underlying oncogenic mechanism under the pathway context based on these multi-dimensional data. CPAD currently documents 4709 entries of experimentally supported relationships between 49 cancers and 1557 paths (some of which corresponding to 41 KEGG pathways) with 2427 upstream regulators across 7 species by reviewing approximately 8500 published papers. What’s more, 33 highly reliable cancer-pathway associations from TCGA projects were integrated. In addition, CPAD also provides predicted upstream regulators for these pathways.
Currently, users can
1. Obtain the detailed descriptions of each experimentally validated cancer-pathway entry;
2. Explore the detailed alteration of genes in cancer pathway at multiple molecular levels of TCGA;
3. Explore the clinical attributes of cancer pathway genes in pan-cancers;
4. Explore drug sensitivity associations of these cancer pathway genes;
5. Explore the experimentally validated or computationally predicted up-stream regulators of cancer pathway
and the detailed information of these regulators at multiple omics level.
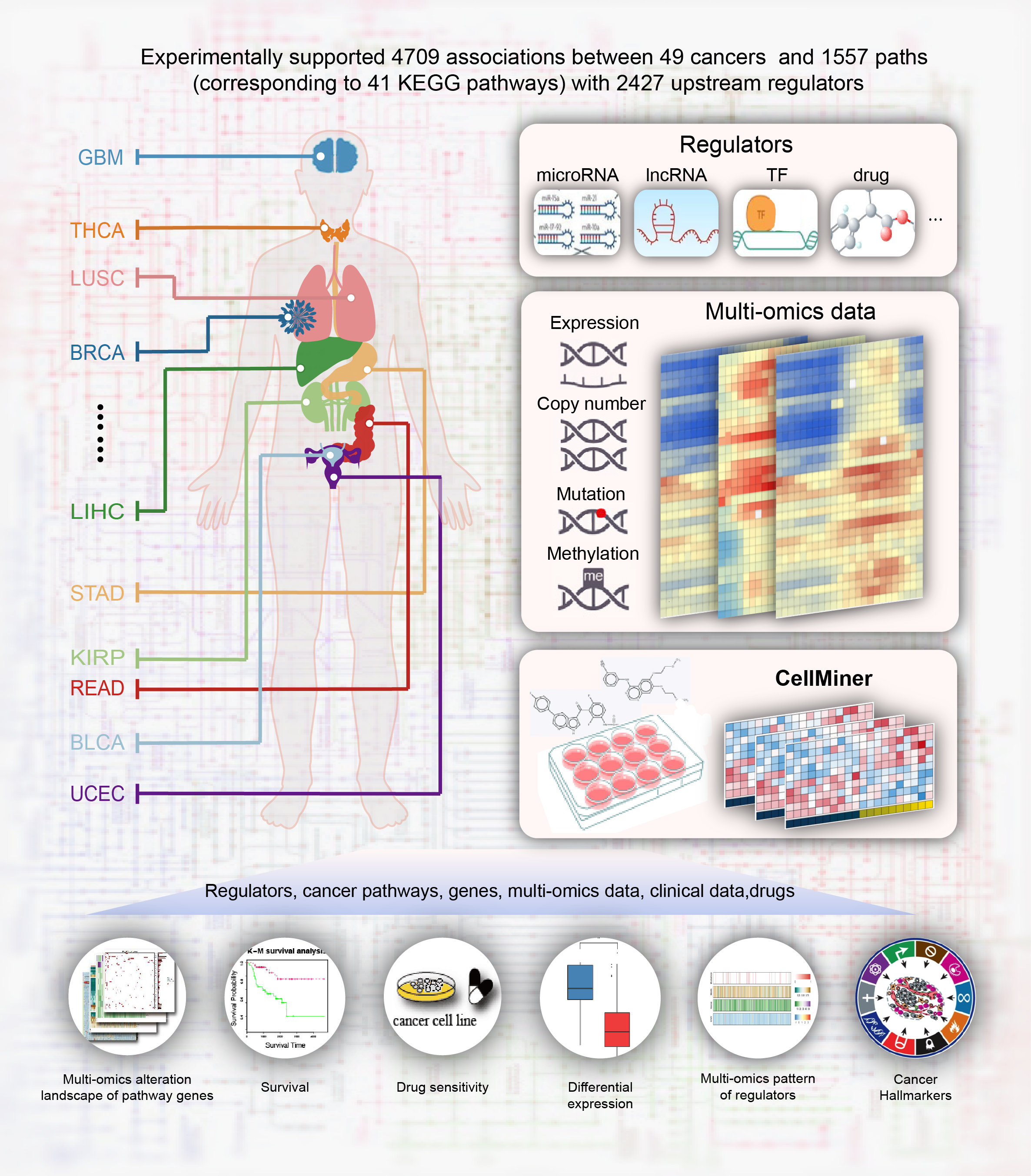
Statistics of experimentally supported relationships
News
CPAD has been formally launched and is freely available at:
http://bio-bigdata.hrbmu.edu.cn/CPAD/. (June 29, 2018)
CPAD has been updated and added 161 associations between 26 cancers and 96 paths from Google Scholar. (February 20, 2019)