1.Database content
Overview of stSNV
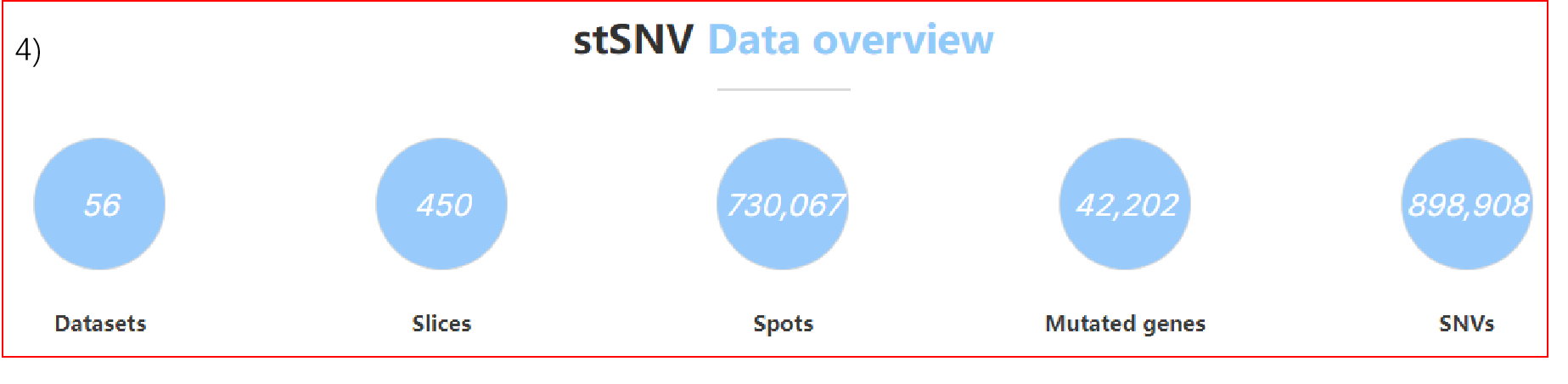
stSNV provides a versatile resource of somatic SNVs (single nucleotide variants) at spatial transcriptomics resolution, involving 21,840 genes in 484,885 spots, covering 27 diseases and 19 normal tissues. Based on a large spatial multicellular dataset, stSNV performs multilevel analyses to provide insights into the spatial variation function in different types of human diseases, and user-friendly comprehensive analysis tools were equipped in the database.
2.Web interface
2.1 stSNV's Home: stSNV provides quick search and highlights
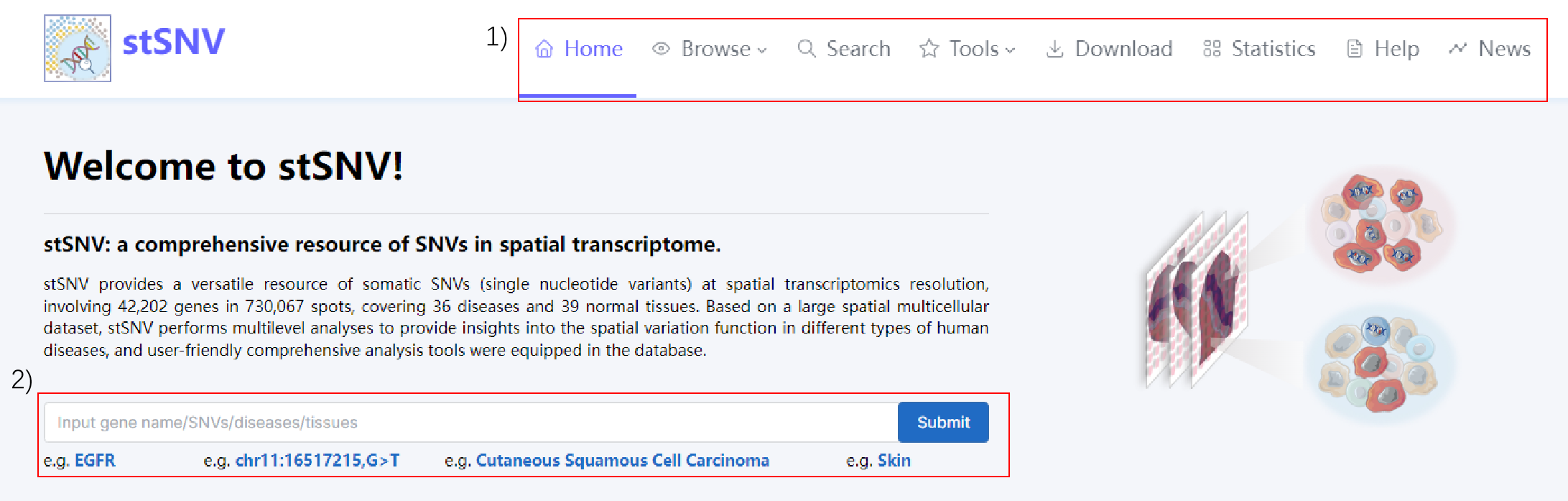
1) stSNV's navigation bar.
2) Enter the name of the gene you are interested in and view its results.
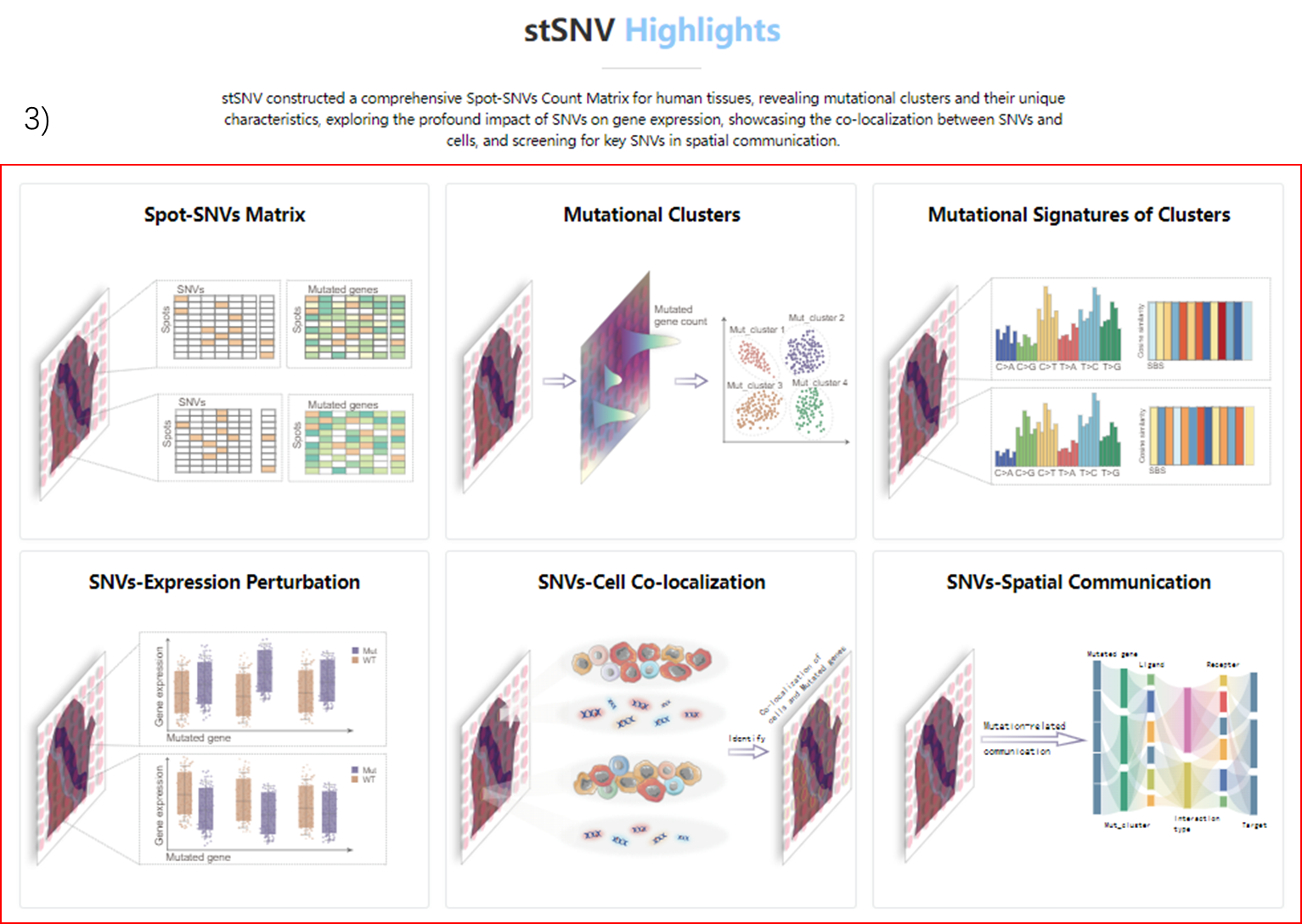
3) View highlights of the database.
4) Data statistics of the database.
2.2 stSNV's Browse: stSNV provides data information of datasets and samples
2.2.1 Datasets
2.2.1.1 Browse result by datasets
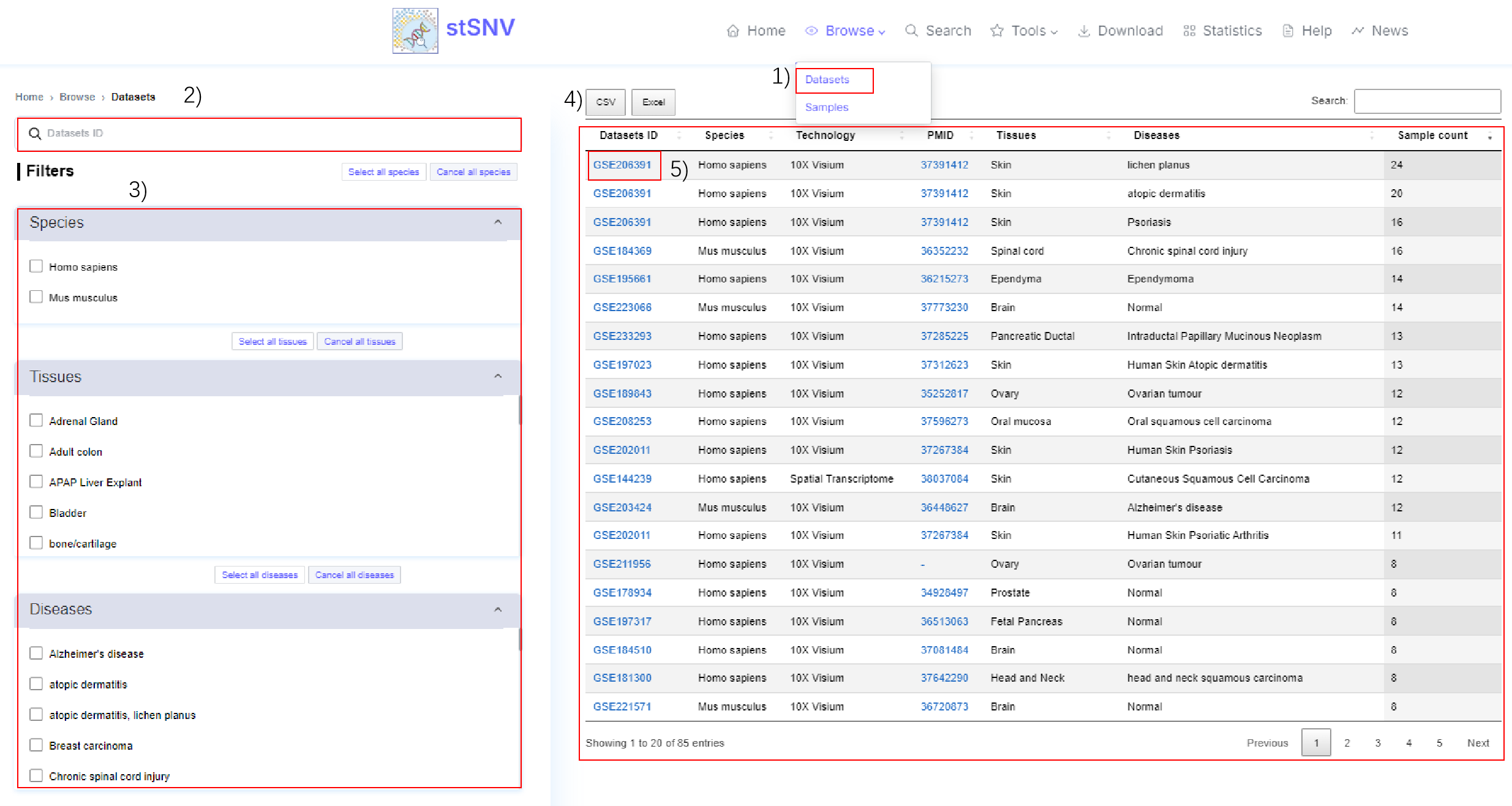
1) Click this button to browse by datasets.
2-3) Global search or filtering datasets.
4) Filtered datasets.
5) Click the Datasets ID to view details.
2.2.1.2 Datasets details
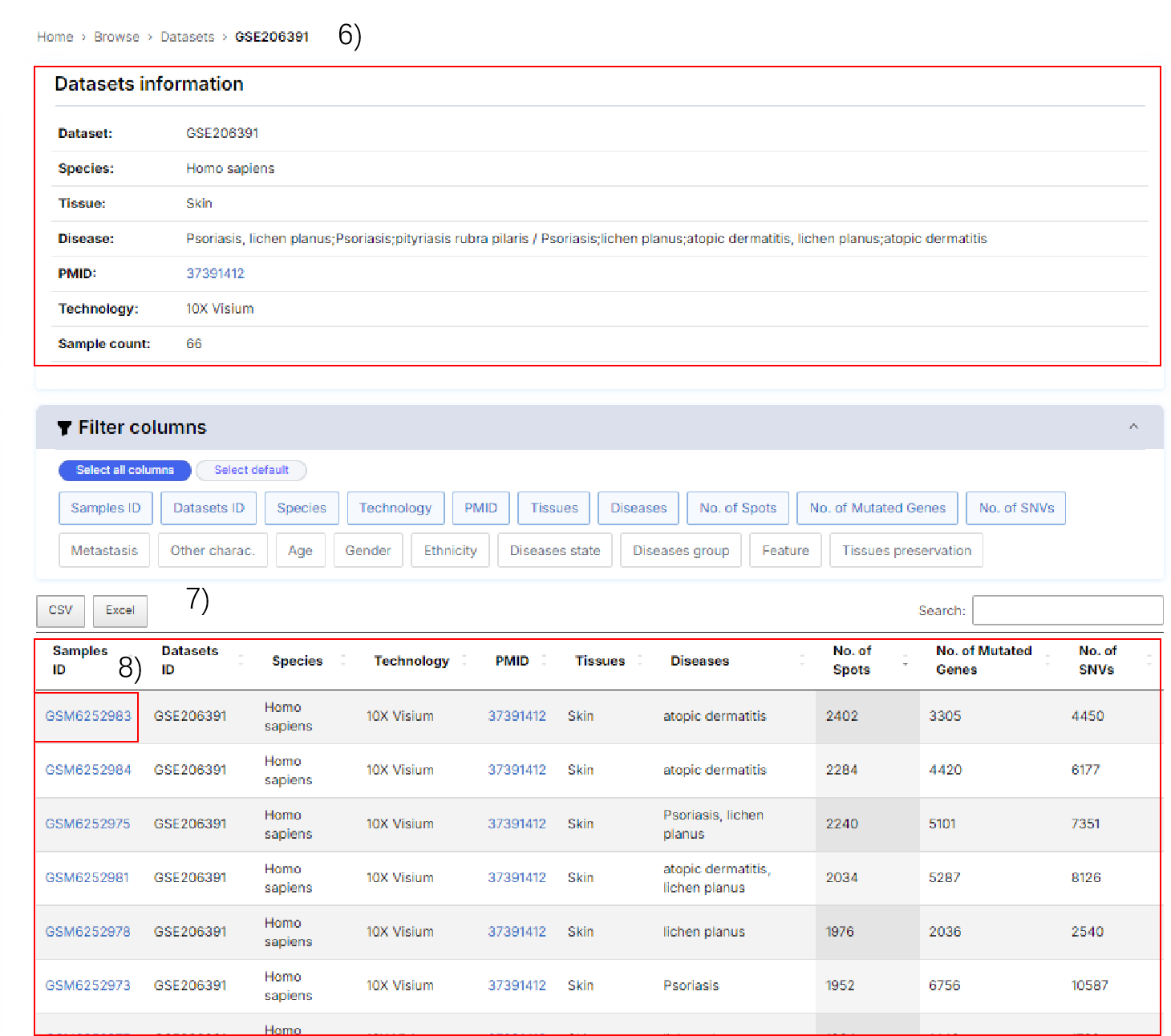
6) Datasets details.
7) Samples included in the dataset.
8) Click the Samples ID to view details.
2.2.2 Browse by samples
2.2.1.1 Browse result by samples
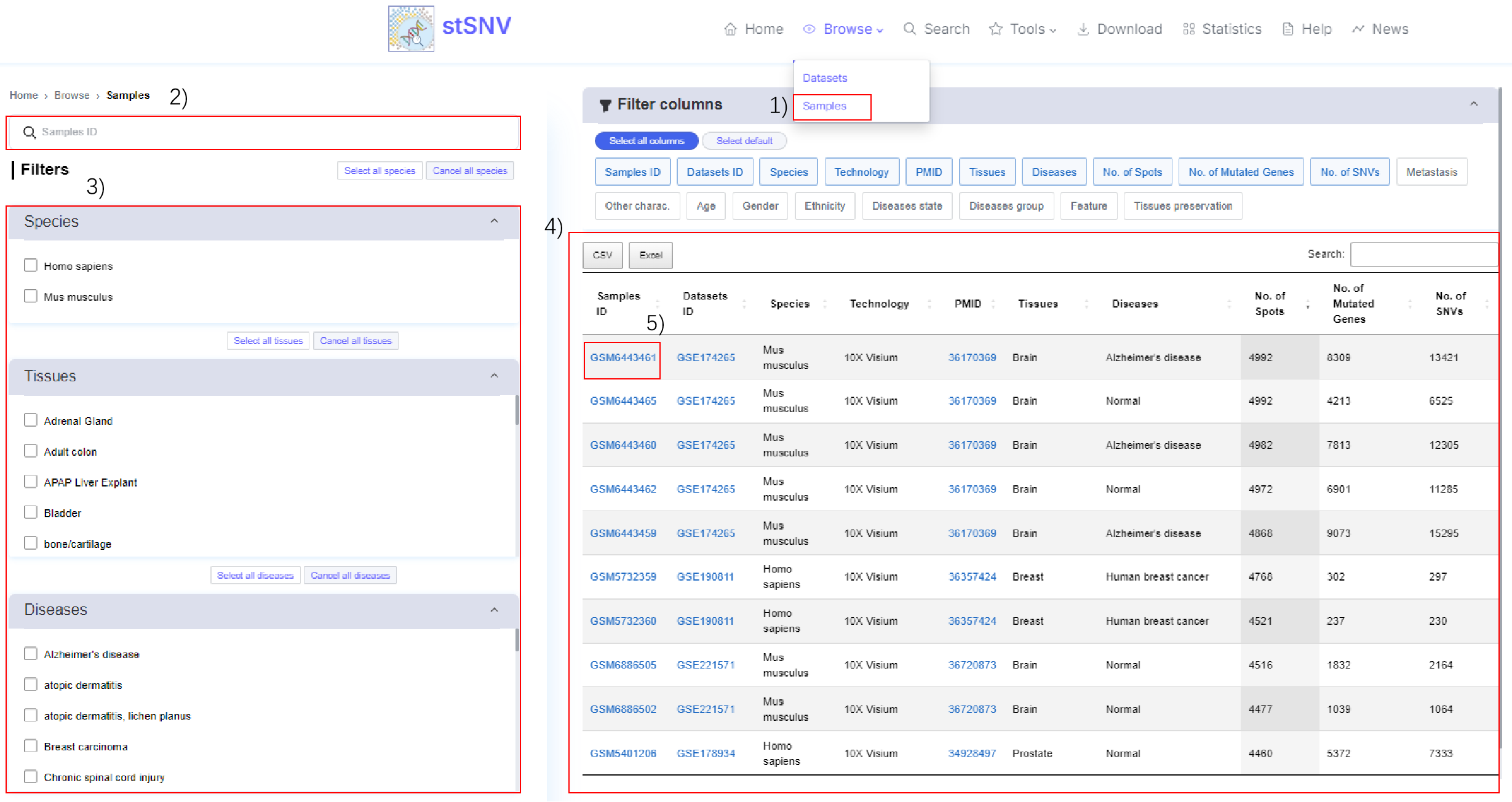
1) Browse - samples.
2-3) Global search or filtering samples.
4) Filtered samples.
5) Click the Samples ID to view the analysis results associated with the sample.
2.2.1.2 Samples details
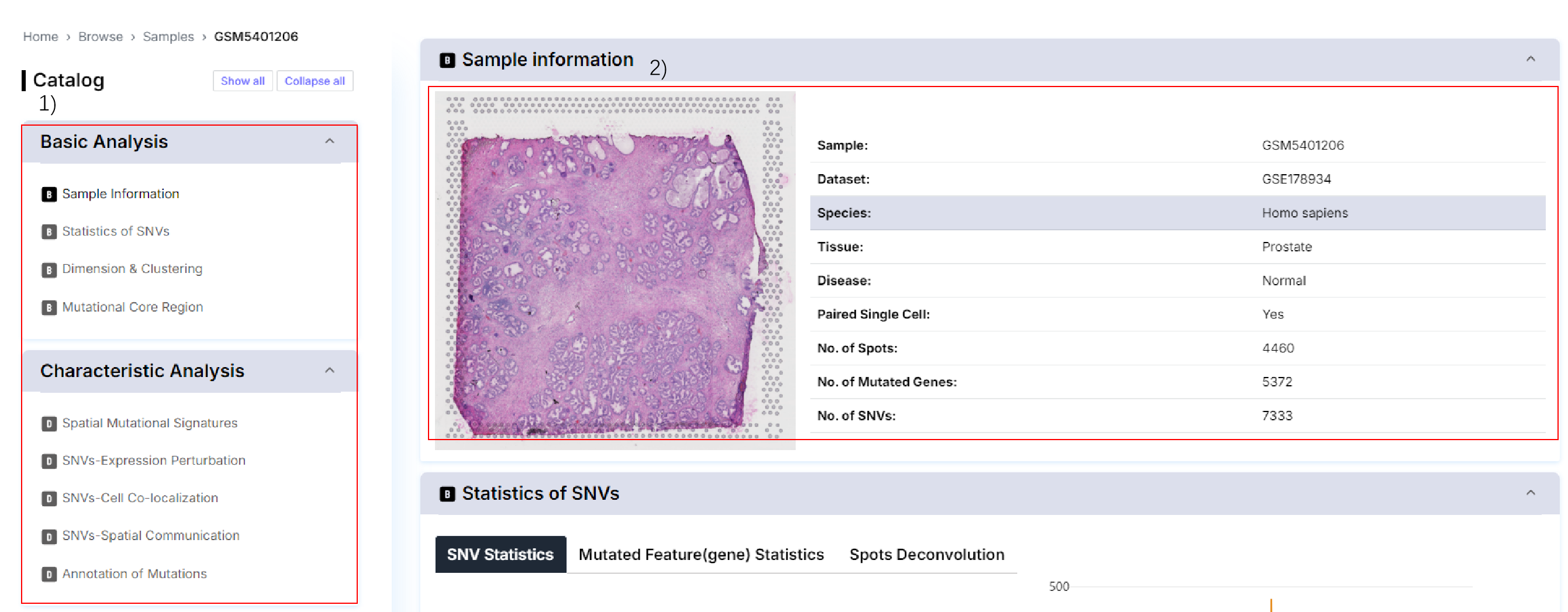
1) Analyze the module directory.
2) Basic information about the current sample.
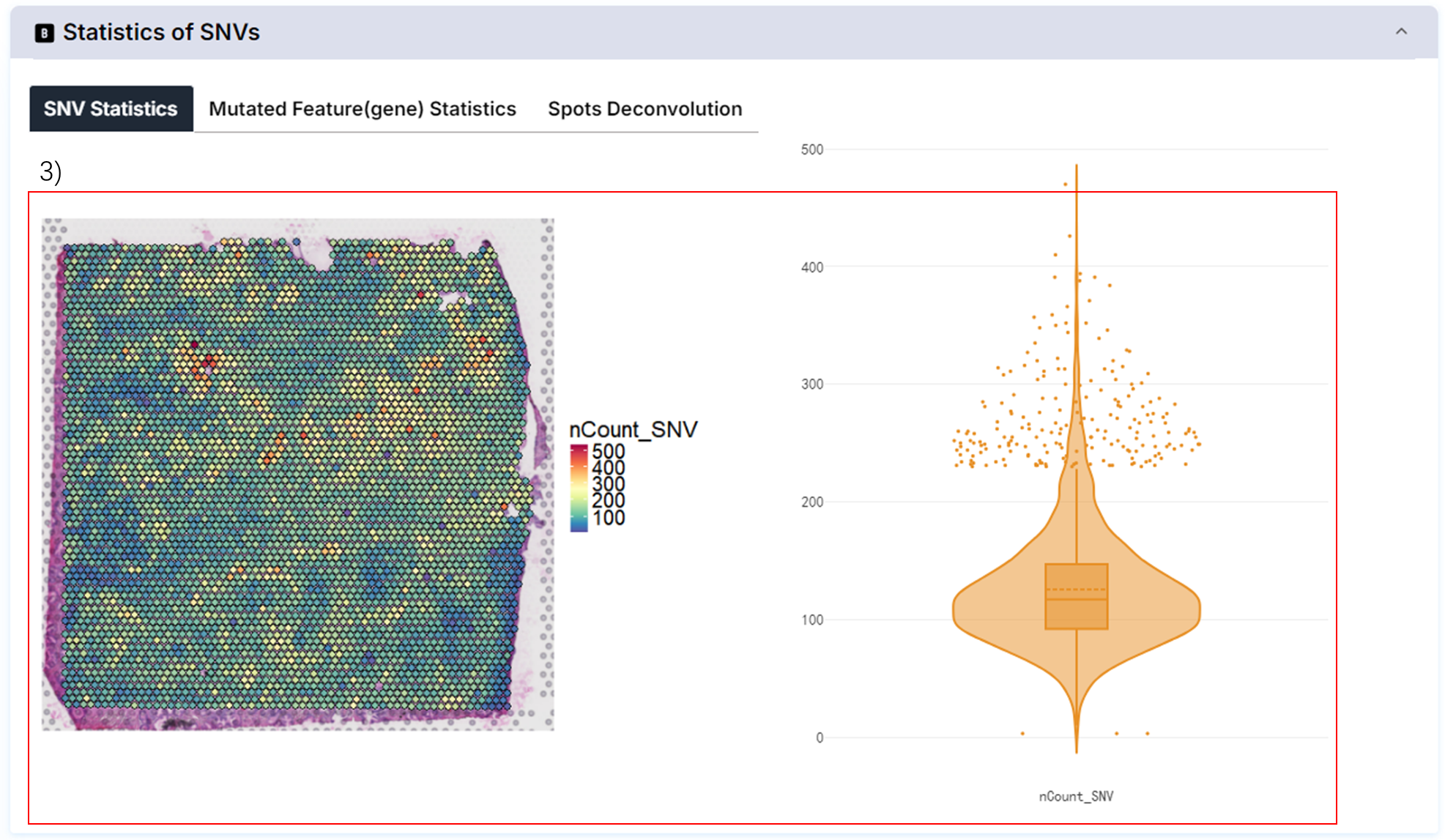
3) The distribution of the number of SNVs identified above the current slice.
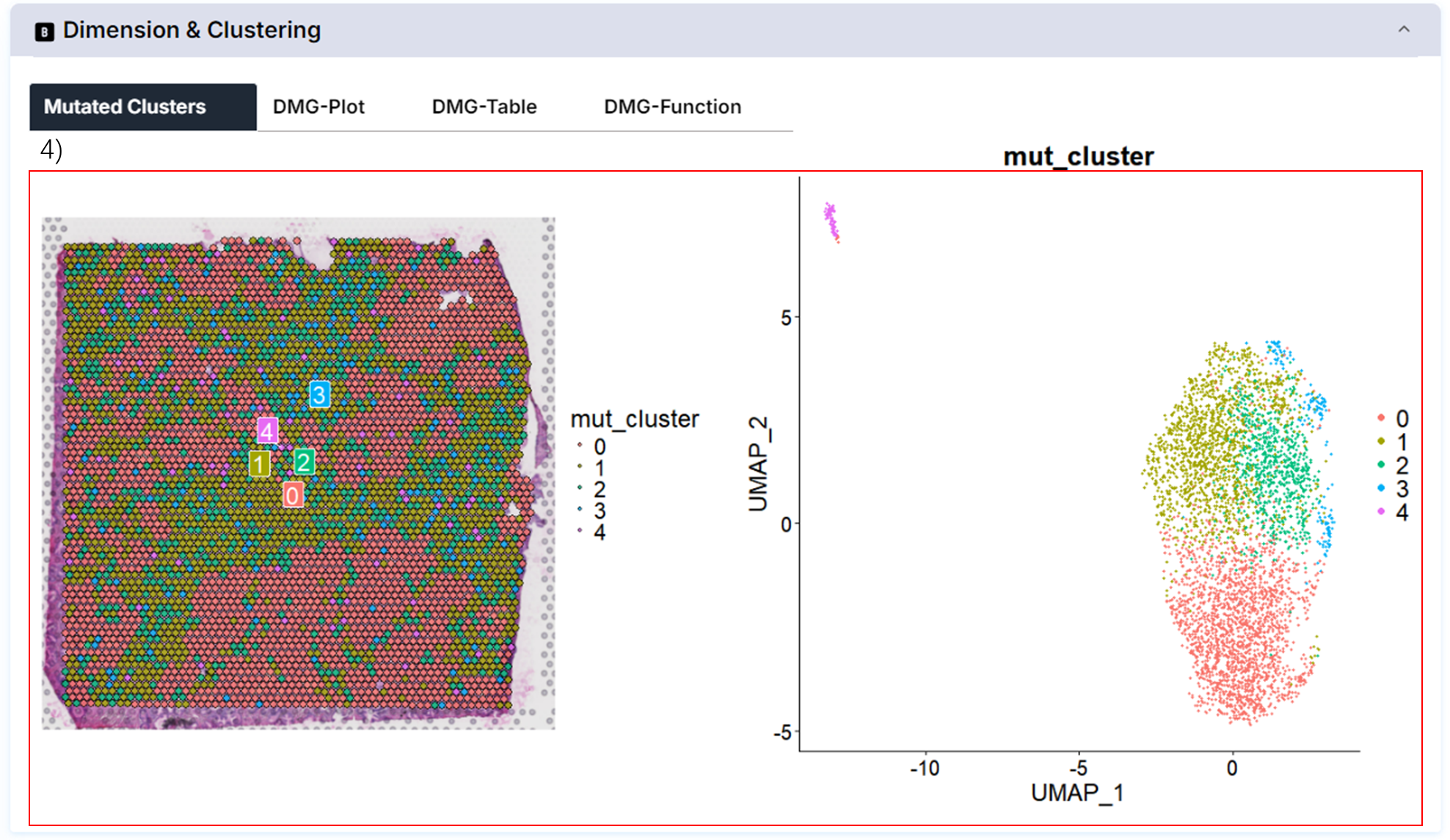
4) Mutational cluster and differential mutational gene.
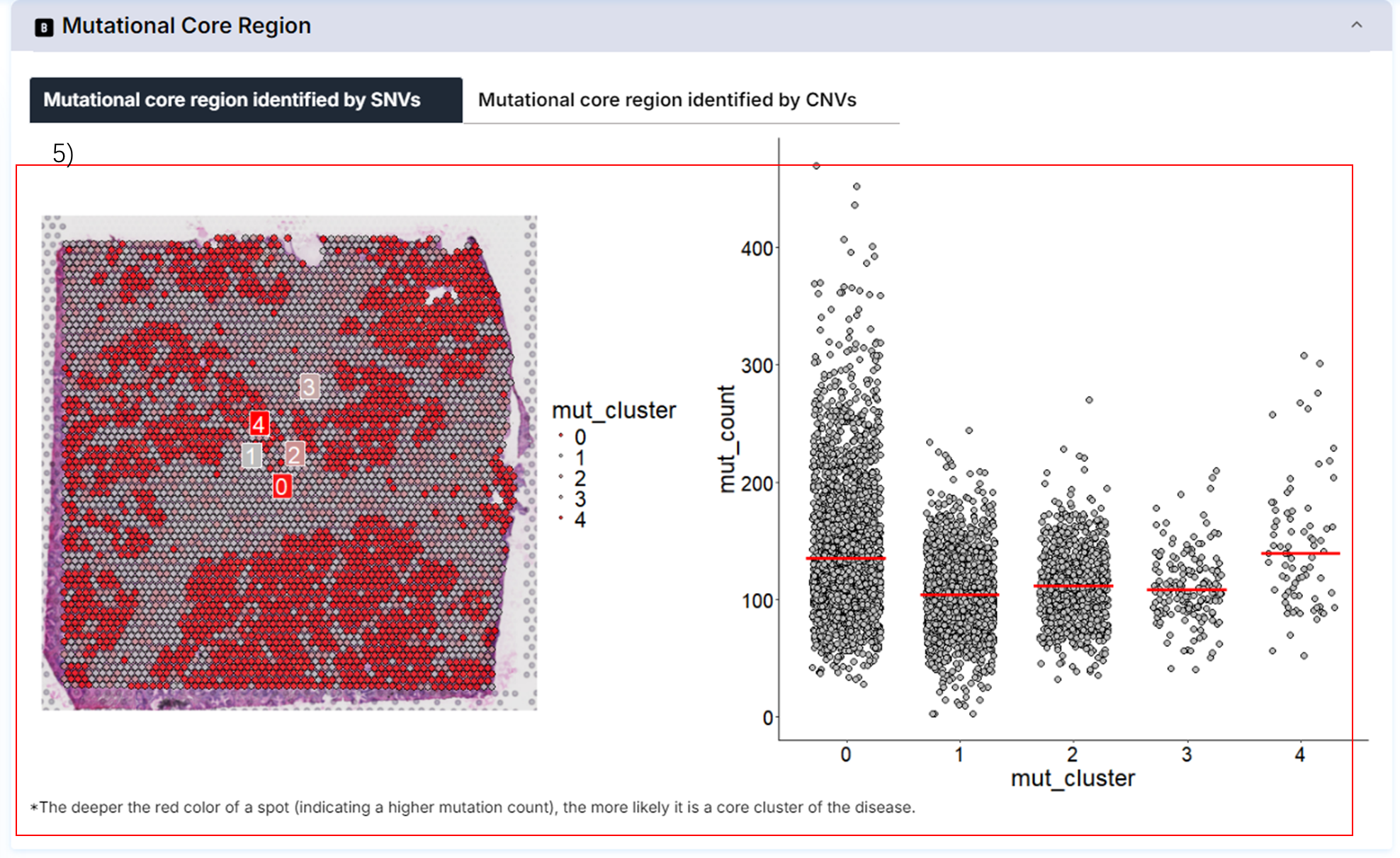
5) Highly variable regions by SNVs and CNVs.
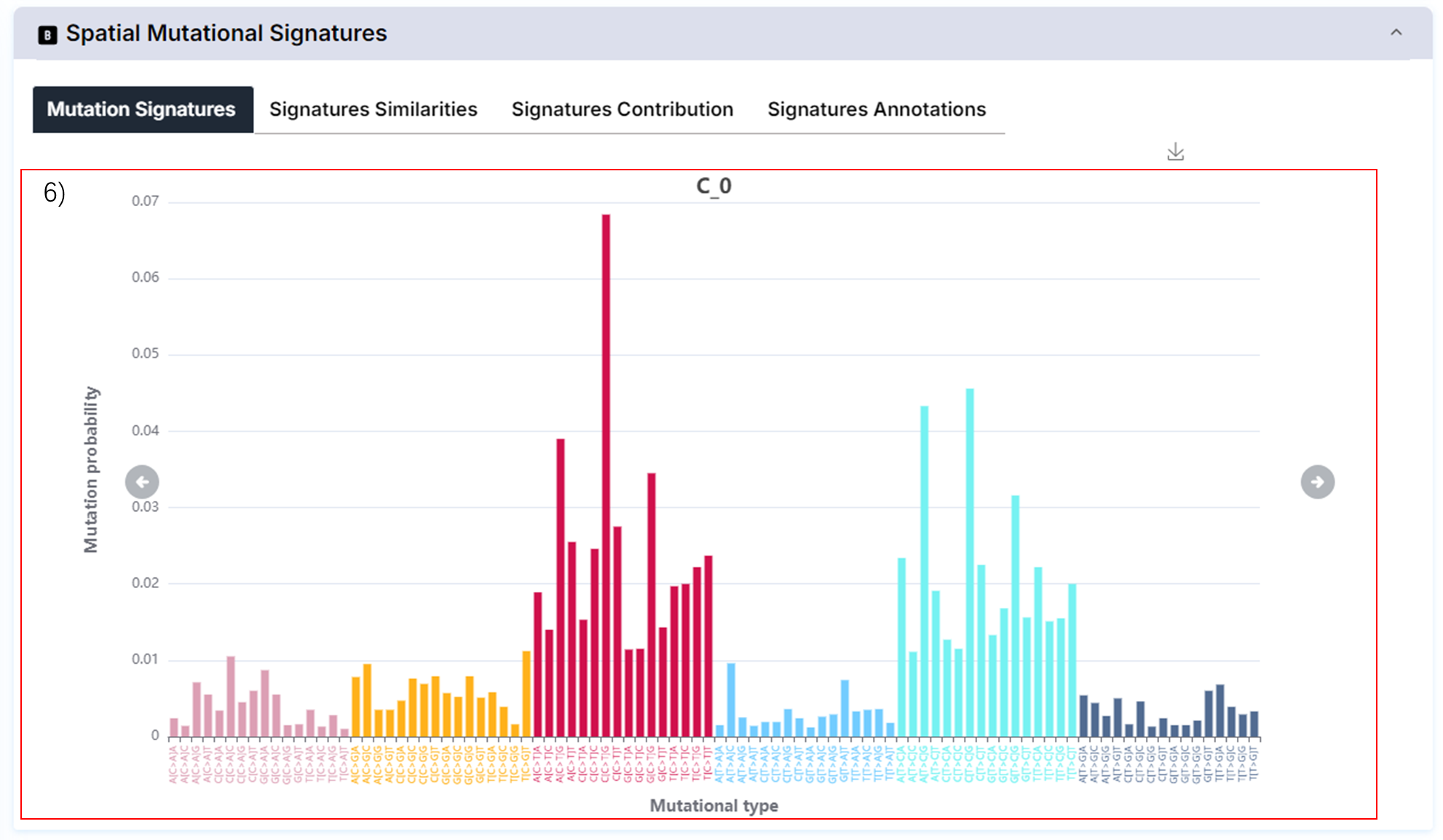
6) The distribution of spatial mutational signatures.
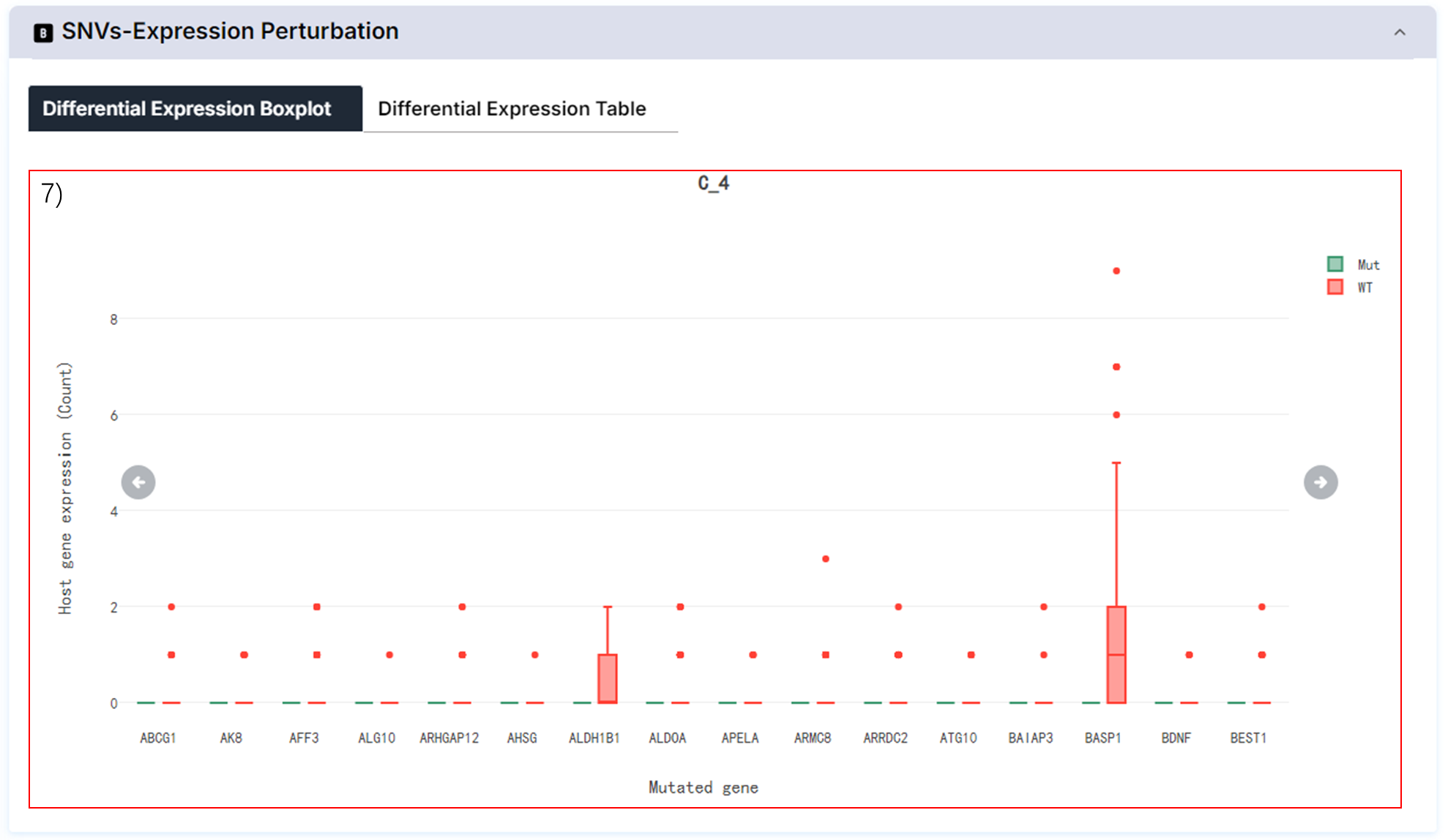
7) Mutated gene perturbed gene expression.
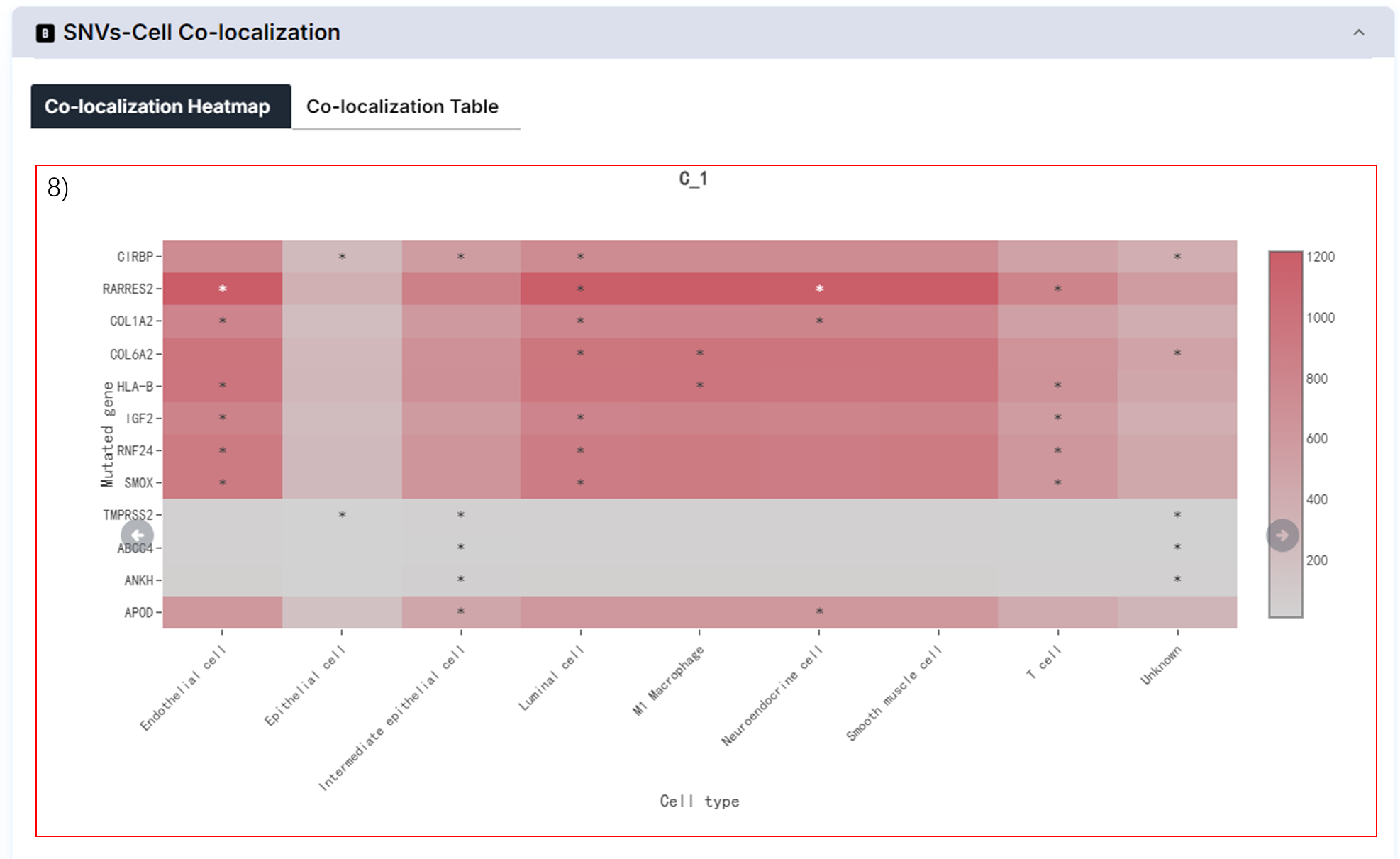
8) Co-localization of mutated genes and cells.
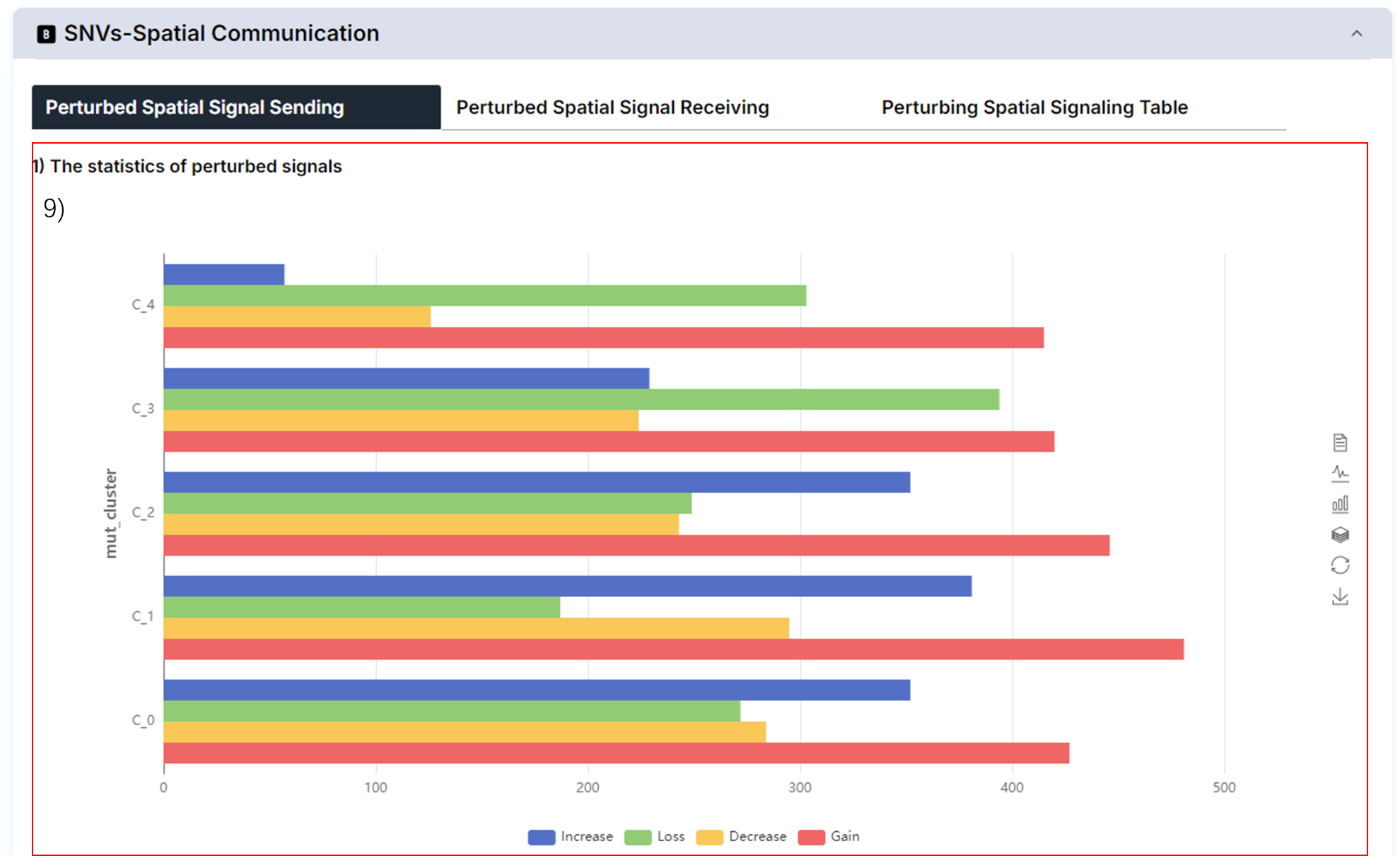
9) Mutated gene perturbed spatial signal sending and receiving.
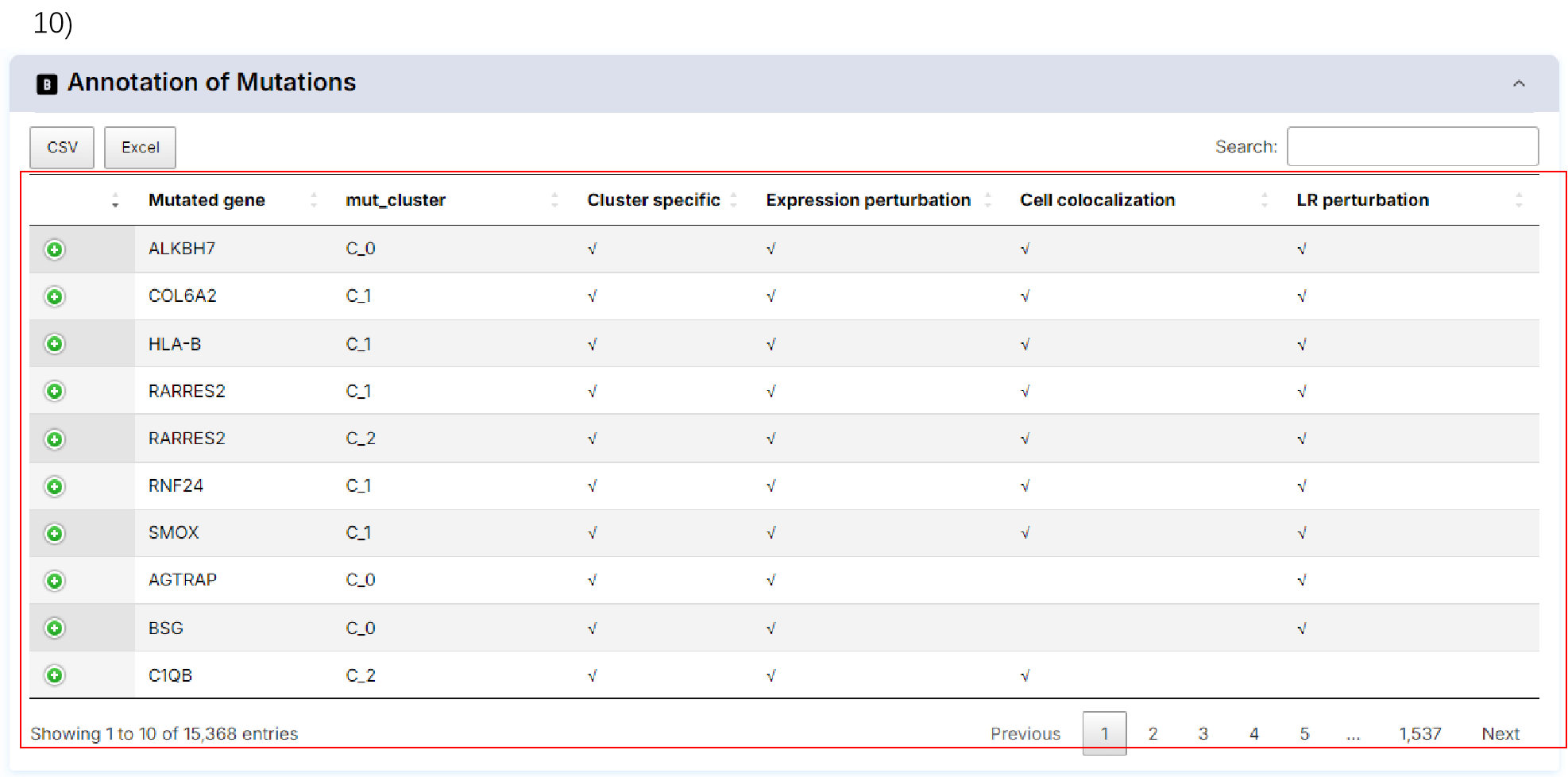
10) Annotation of mutations.
2.3 stSNV's Search: stSNV provides a user-friendly search and result interface
2.3.1 Search results
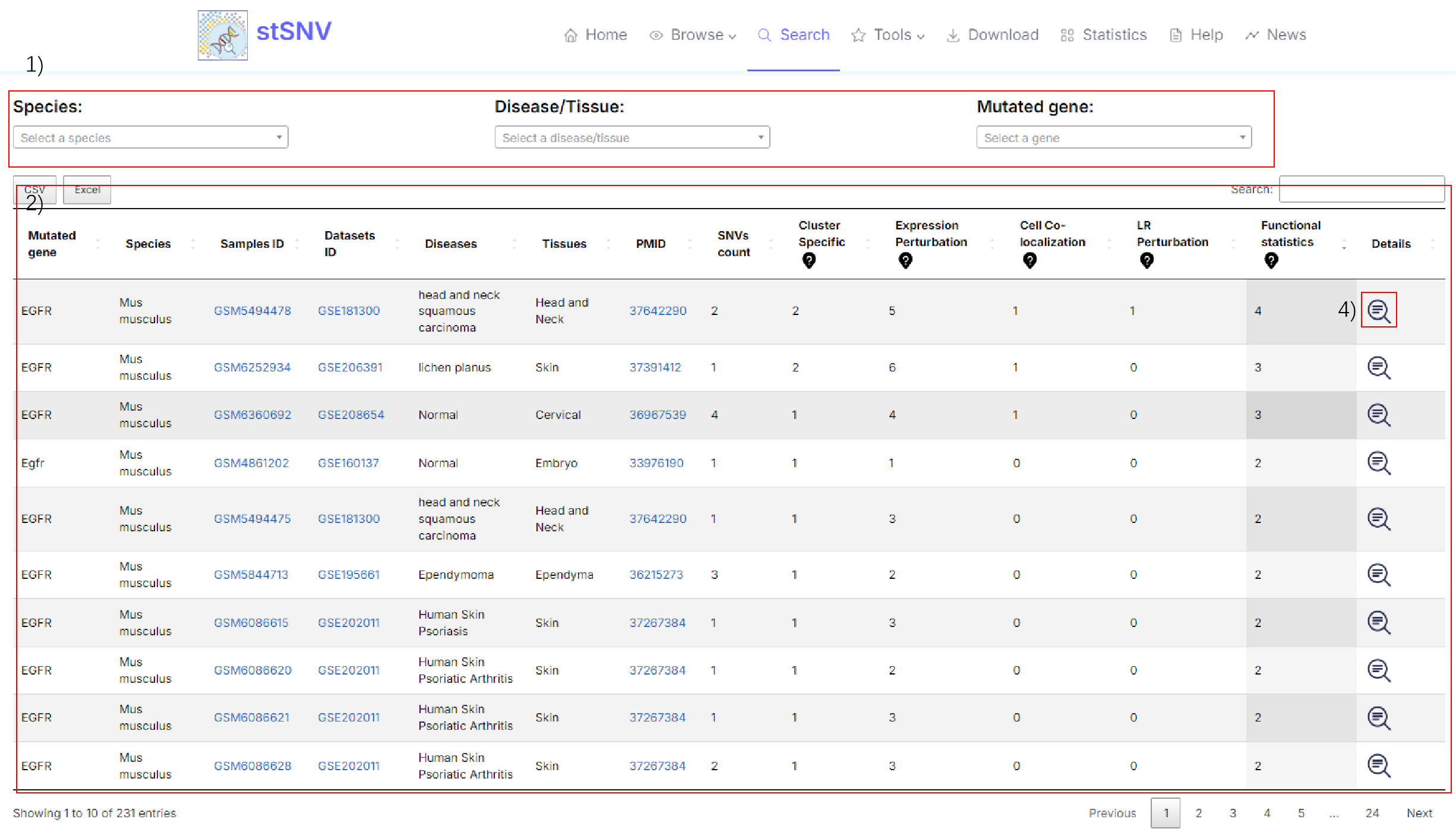
1) Search for mutated genes of interest.
2) Results of mutated gene analysis in samples.
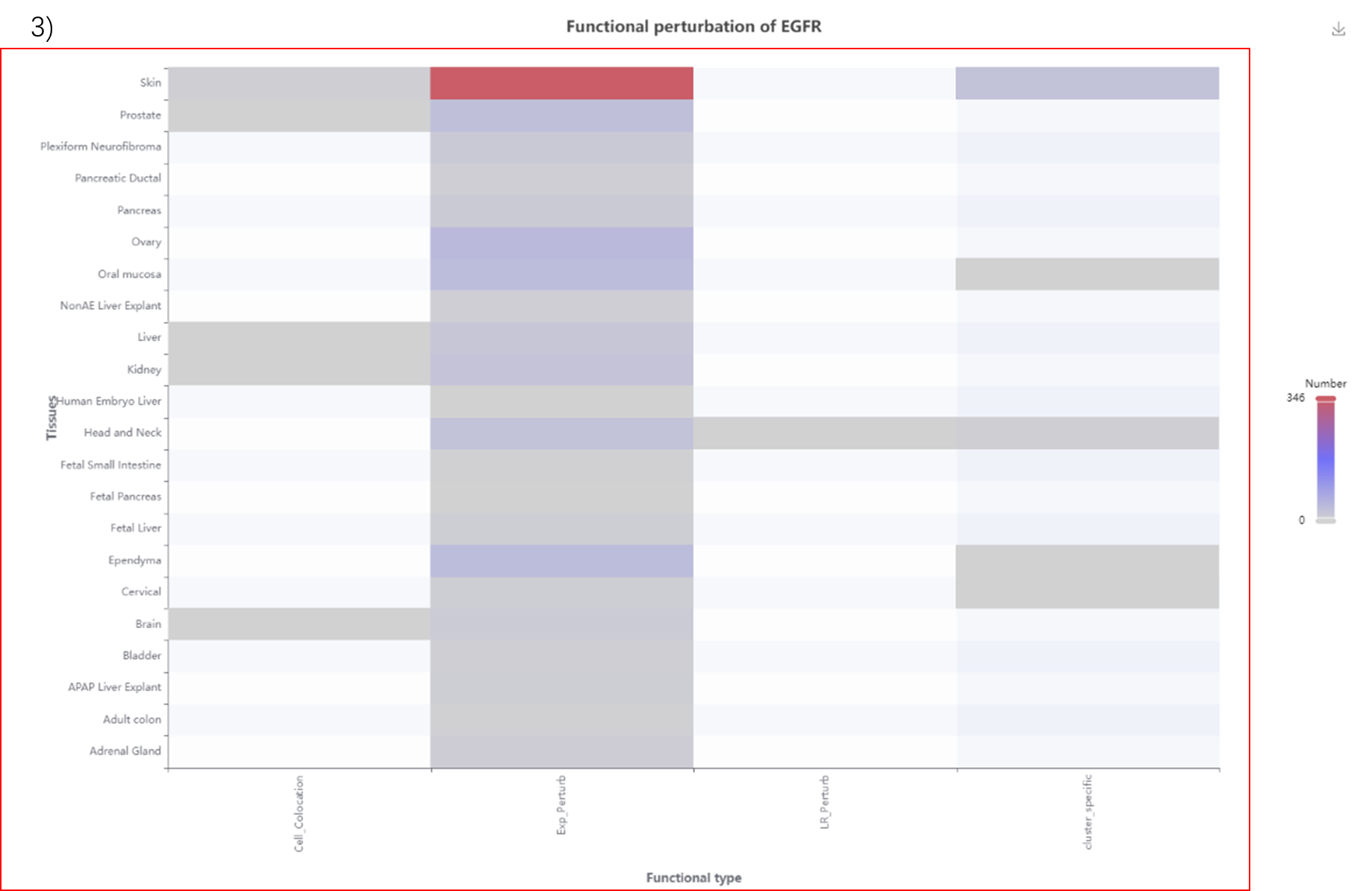
3) Heatmap of mutated gene analysis in samples.
4) Click to see details of the functionally mutated gene in the spacial sample.
2.3.2 Search details
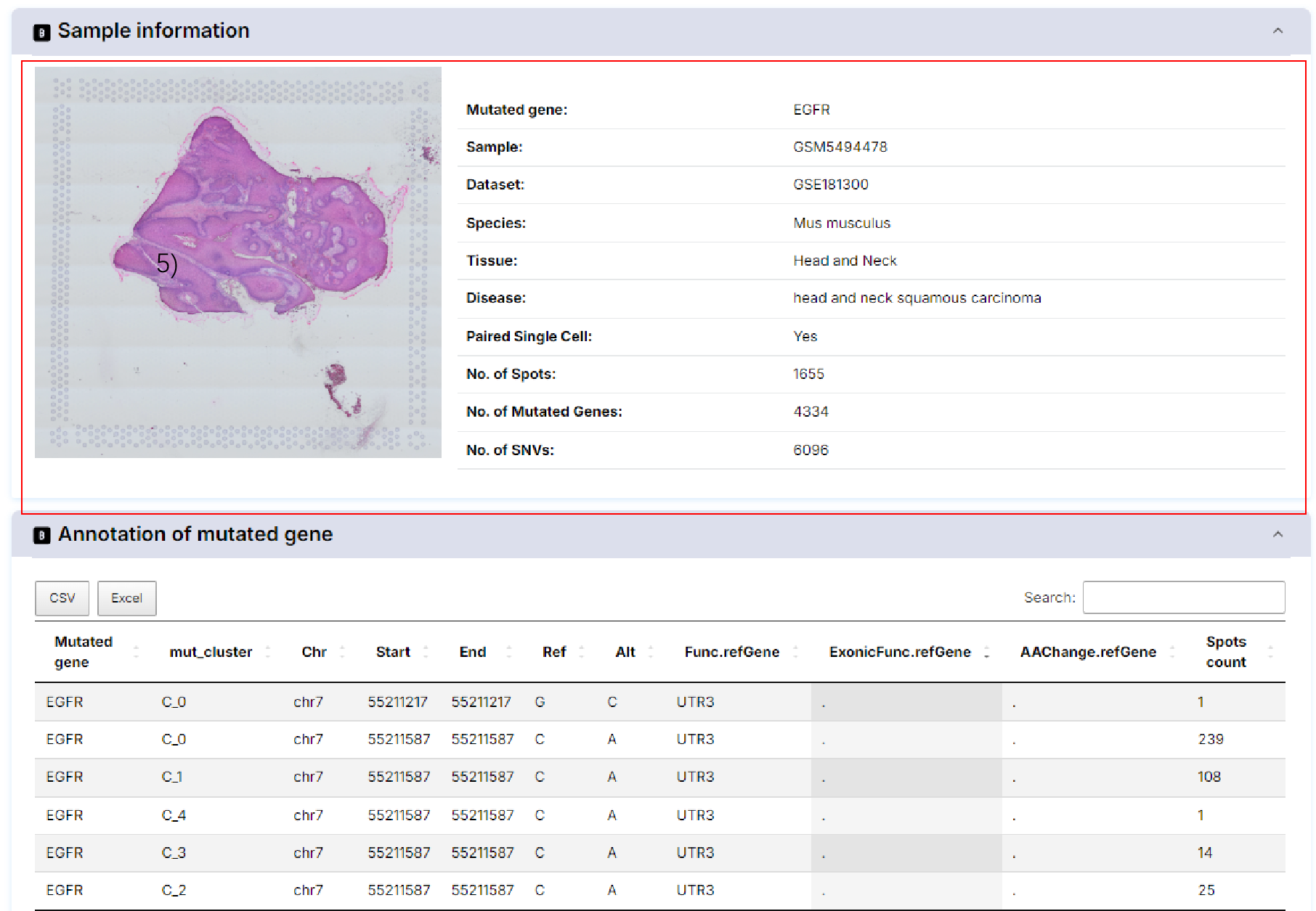
5) Functions of mutated gene.
2.4 stSNV's Tools: stSNV provides 2 user-friendly tools
2.4.1 Tools-Spatial colocalization
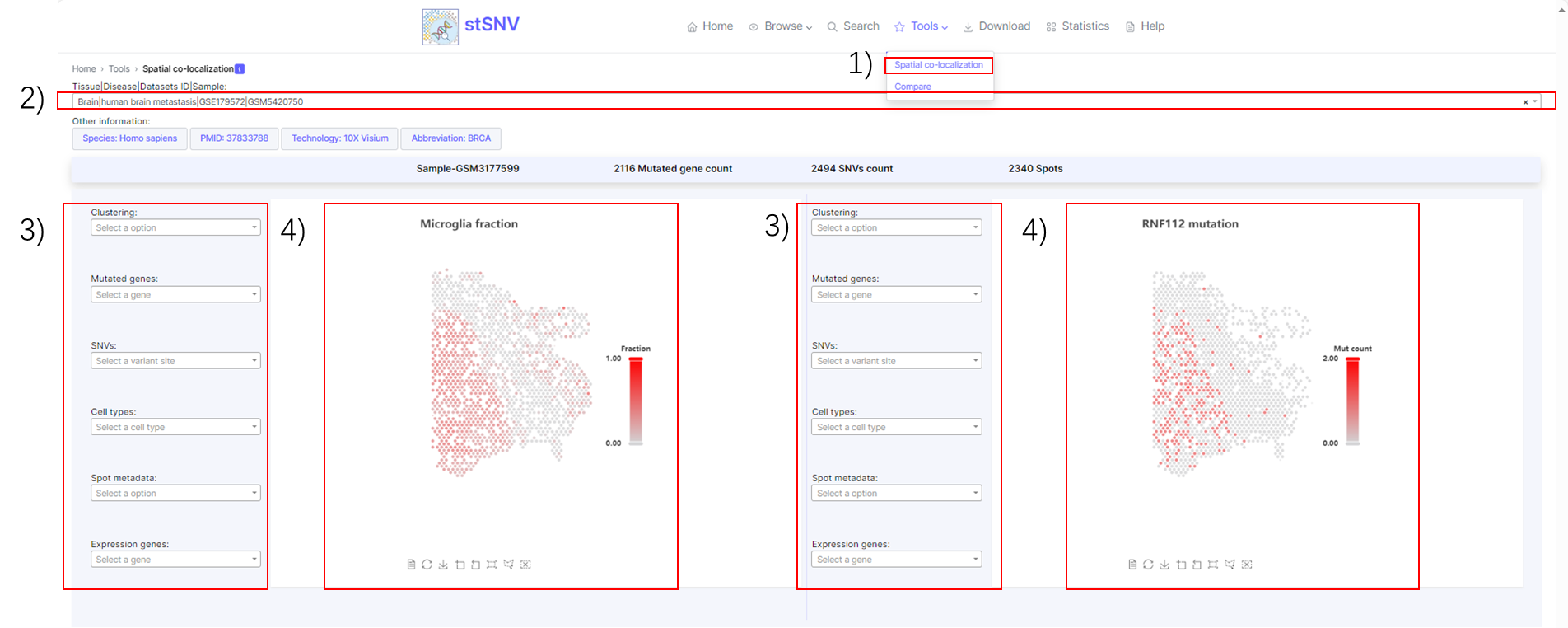
1) Tools- Spatial co-localization.
2) Search sample.
3) Filter spatial features of interest such as mutational cluster, mutated gene, cell type.
4) Spatial features visualization.
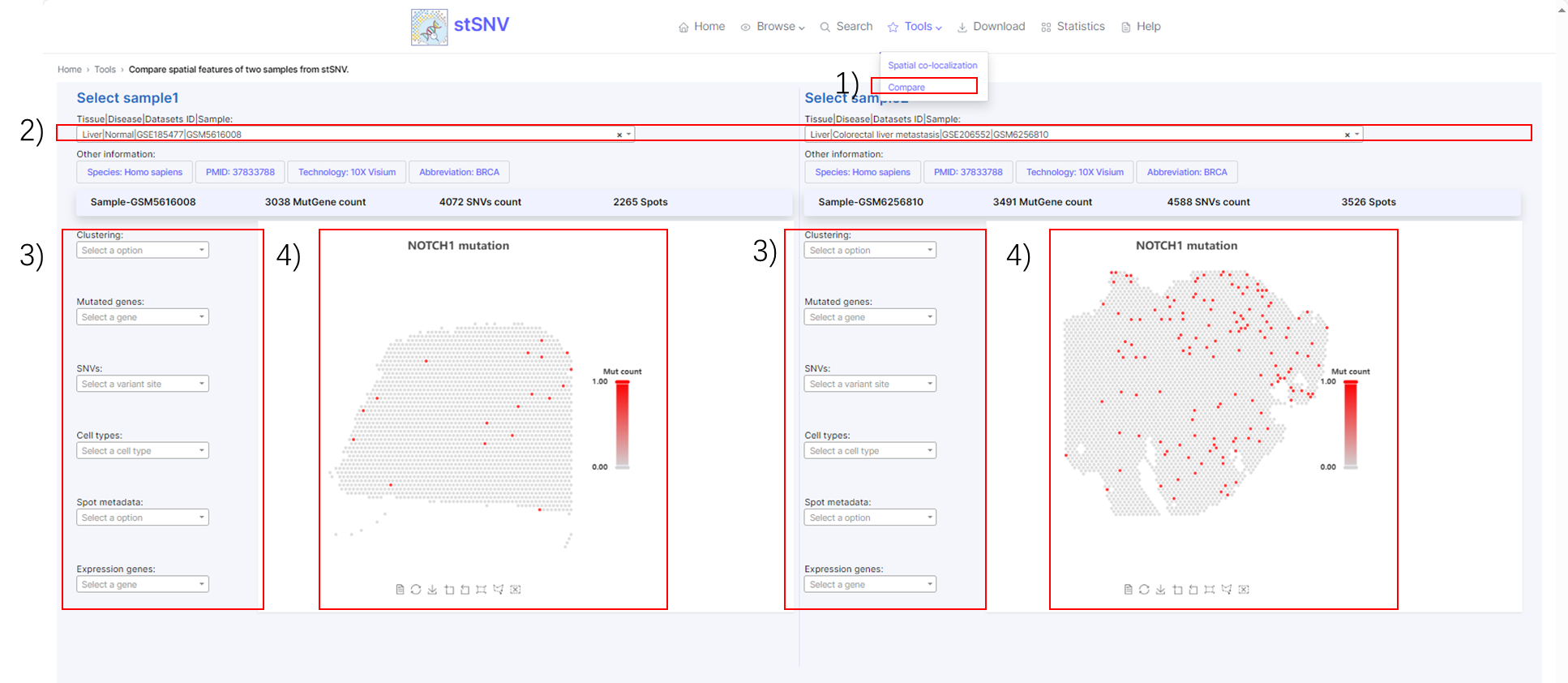
2.4.2 Tools-Compare
1) Tools-Compare.
2) Search sample.
3) Filter spatial features of interest such as mutational cluster, mutated gene, cell type.
4) Spatial features visualization.
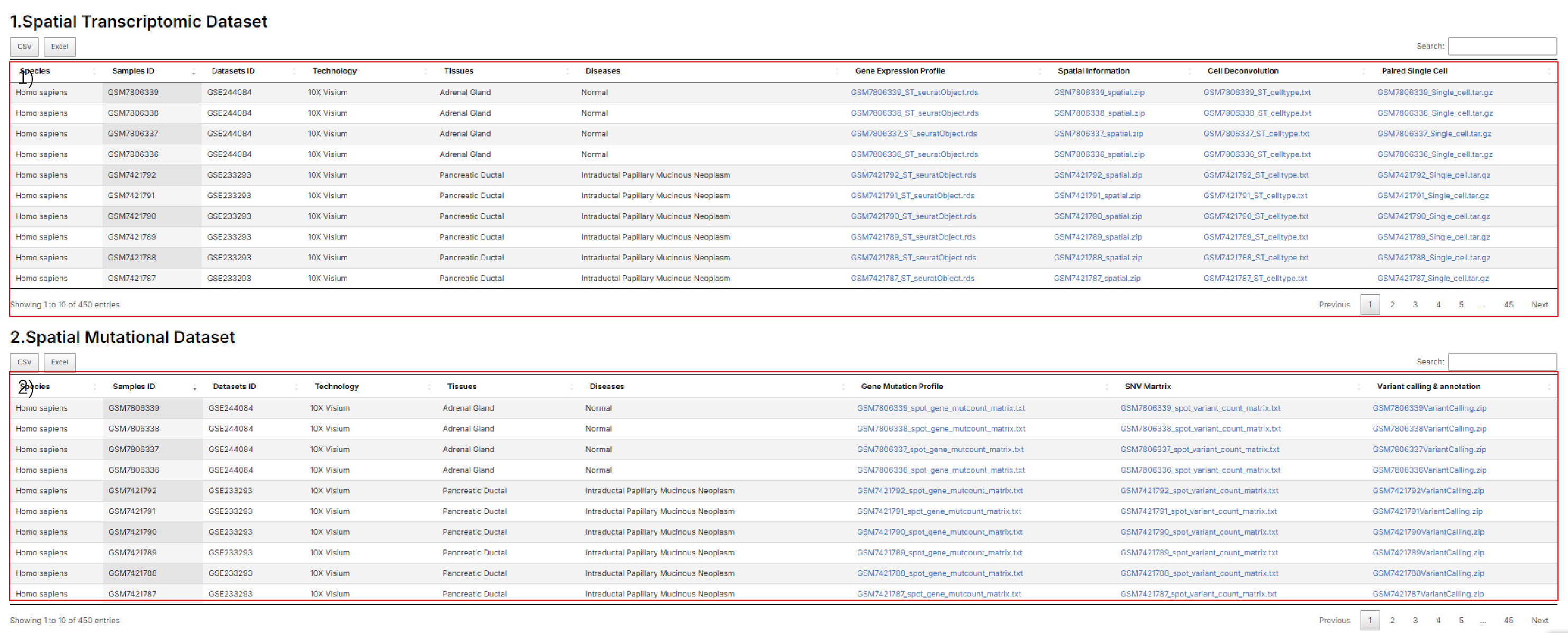
2.5 stSNV's Download
1) Spatial transcriptomics. 2) Spatial Mutational genomics.
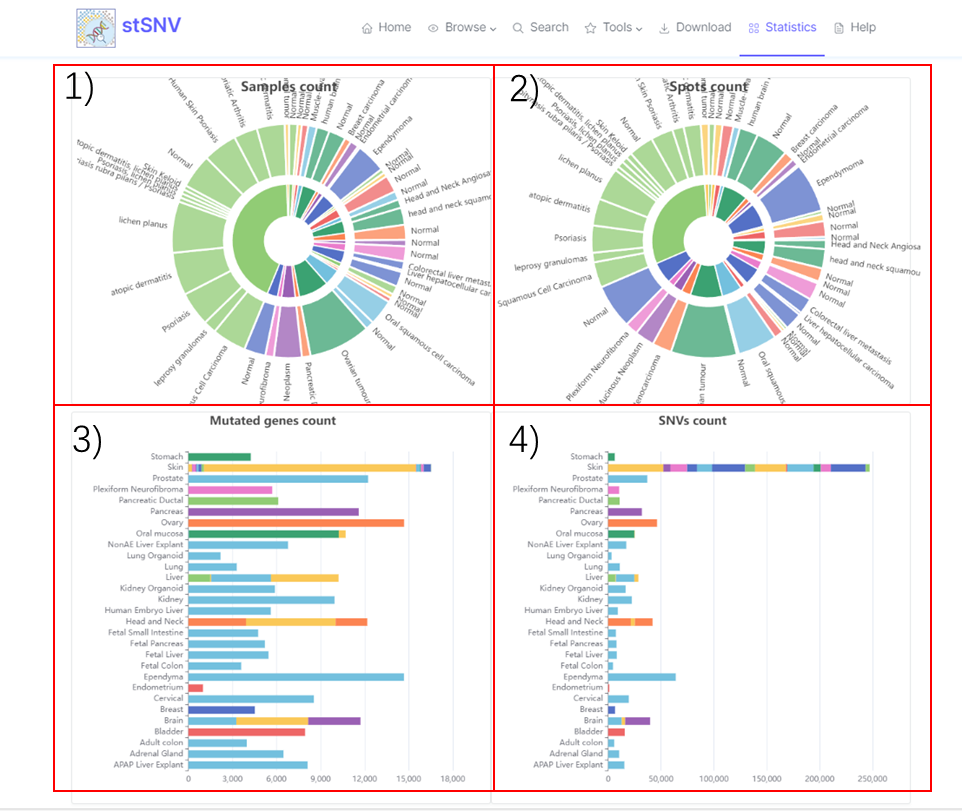
2.6 stSNV's Statistics
1) Mutational sample statistics. 2) Mutational spot statistics. 3) Mutational gene statistics. 4) SNV statistics.